Molecular Recognition as an Information Channel: The Role of Conformational Changes
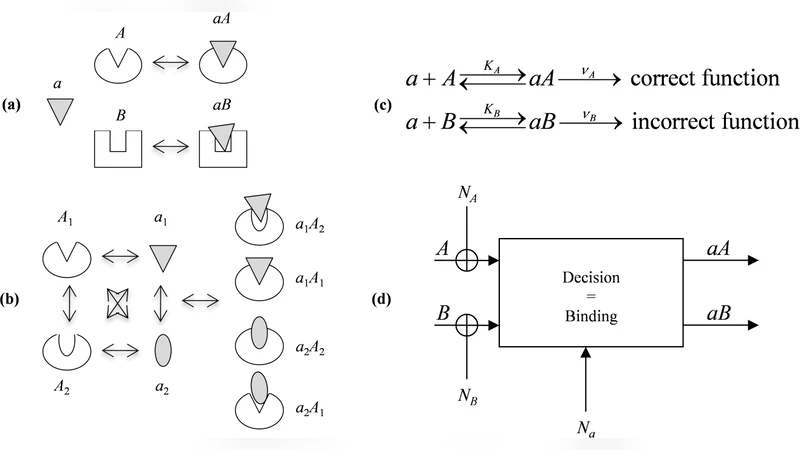
Molecular recognition, which is essential in processing information in biological systems, takes place in a crowded noisy biochemical environment and requires the recognition of a specific target within a background of various similar competing molecules. We consider molecular recognition as a transmission of information via a noisy channel and use this analogy to gain insights on the optimal, or fittest, molecular recognizer. We focus on the optimal structural properties of the molecules such as flexibility and conformation. We show that conformational changes upon binding, which often occur during molecular recognition, may optimize the detection performance of the recognizer. We thus suggest a generic design principle termed ‘conformational proofreading’ in which deformation enhances detection. We evaluate the optimal flexibility of the molecular recognizer, which is analogous to the stochasticity in a decision unit. In some scenarios, a flexible recognizer, i.e., a stochastic decision unit, performs better than a rigid, deterministic one. As a biological example, we discuss conformational changes during homologous recombination, the process of genetic exchange between two DNA strands.
💡 Research Summary
The paper reframes molecular recognition as a communication problem in which a recognizer (the transmitter) must extract a specific target signal from a noisy biochemical channel populated by many similar, competing molecules. By mapping binding events onto information‑theoretic quantities, the authors treat the recognizer’s structural properties—particularly flexibility and the ability to undergo conformational changes—as parameters that determine channel capacity and detection performance.
A central insight is that a recognizer’s rigidity versus flexibility plays a role analogous to deterministic versus stochastic decision units in signal processing. A rigid molecule presents a fixed binding geometry, which can be efficient when the signal‑to‑noise ratio (the energetic distinction between correct and incorrect partners) is high. However, in crowded environments where non‑target molecules are energetically similar, a certain degree of flexibility allows the recognizer to explore multiple conformations, thereby increasing the probability of finding a low‑energy, highly specific binding mode for the true target while raising the energetic barrier for spurious interactions.
The authors formalize this trade‑off through a “conformational proofreading” principle. Upon initial contact, the recognizer deforms; only if the partner is the correct target does the deformation lead to a stable, low‑energy complex. Incorrect partners either fail to induce the proper conformational shift or are left in a high‑energy strained state, effectively filtering them out. Mathematically, the model combines Bayes decision theory with elastic energy terms (characterized by a stiffness constant) and binding free‑energy differences (ΔG). Optimization of these parameters shows that, for a given noise level, there exists an optimal stiffness that maximizes the probability of correct detection. In many realistic scenarios—especially when the energetic gap between target and competitors is small—a flexible, stochastic recognizer outperforms a rigid, deterministic one.
The biological illustration focuses on homologous recombination mediated by RecA. RecA binding stretches and bends DNA, creating a conformational checkpoint that preferentially stabilizes correctly aligned homologous sequences while destabilizing mismatched ones. This real‑world example demonstrates how nature exploits conformational proofreading to achieve high fidelity in genetic exchange.
Overall, the study proposes a generic design rule for both natural and engineered molecular sensors: incorporate controlled conformational changes to act as a proofreading step, thereby enhancing specificity without sacrificing sensitivity. The framework bridges molecular biophysics with information theory and suggests new avenues for designing flexible biosensors, drug molecules, and synthetic biological circuits that operate optimally in noisy cellular environments.
Comments & Academic Discussion
Loading comments...
Leave a Comment