StochKit-FF: Efficient Systems Biology on Multicore Architectures
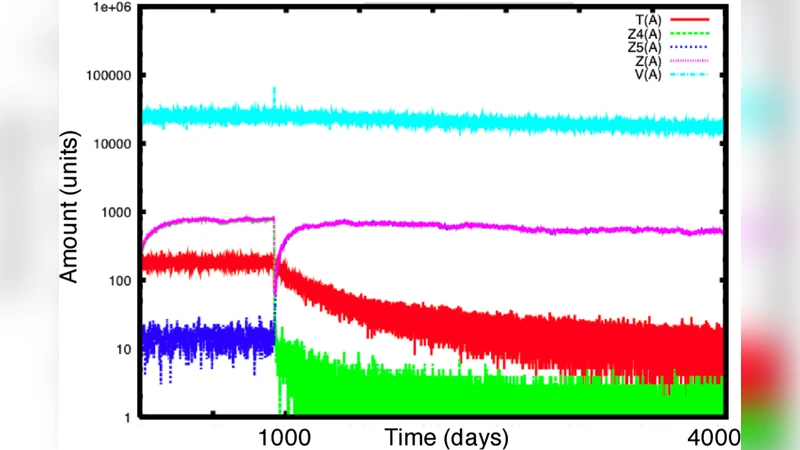
The stochastic modelling of biological systems is an informative, and in some cases, very adequate technique, which may however result in being more expensive than other modelling approaches, such as differential equations. We present StochKit-FF, a parallel version of StochKit, a reference toolkit for stochastic simulations. StochKit-FF is based on the FastFlow programming toolkit for multicores and exploits the novel concept of selective memory. We experiment StochKit-FF on a model of HIV infection dynamics, with the aim of extracting information from efficiently run experiments, here in terms of average and variance and, on a longer term, of more structured data.
💡 Research Summary
The paper introduces StochKit‑FF, a parallel extension of the widely used stochastic simulation toolkit StochKit, designed to exploit modern multicore processors for large‑scale systems biology experiments. The authors begin by highlighting the inherent computational expense of exact stochastic simulation algorithms (SSAs), such as Gillespie’s direct method, especially when thousands or millions of independent trajectories are required for statistical inference. While StochKit already offers a high‑quality reference implementation, its original design targets single‑core execution, leaving a performance gap on today’s many‑core hardware.
To bridge this gap, the authors build StochKit‑FF on top of the FastFlow programming framework, which supplies a lock‑free, cache‑friendly pipeline model and a “farm” pattern for embarrassingly parallel workloads. The key innovation is the concept of selective memory, a runtime aggregation mechanism that reduces the volume of data transferred between worker threads and the collector thread. Each worker thread runs an independent SSA instance, storing intermediate state vectors locally. Instead of writing every trajectory to disk, the selective memory module maintains per‑time‑window aggregates (sum and sum‑of‑squares) that enable on‑the‑fly computation of means, variances, and higher‑order moments. This approach dramatically cuts I/O bandwidth, memory footprint, and synchronization overhead.
Implementation details include: (1) wrapping the original StochKit C++ core in FastFlow’s API; (2) using thread‑local random number generators to avoid contention; (3) employing a sliding‑window algorithm that updates statistical accumulators in O(1) time per event; and (4) configuring the pipeline as a three‑stage flow—workers → selective memory → output—so that the collector can run on a dedicated core without becoming a bottleneck. The authors also discuss how FastFlow’s lock‑free queues guarantee near‑linear scalability up to the number of physical cores, provided the workload remains embarrassingly parallel.
The experimental evaluation focuses on two case studies. The primary benchmark is an HIV infection dynamics model comprising twelve reactions (cell infection, viral production, immune response, etc.). The authors execute 10 000 independent trajectories on a 16‑core Intel Xeon system, comparing three configurations: (a) original StochKit on a single core, (b) StochKit‑FF without selective memory (full trajectory logging), and (c) StochKit‑FF with selective memory. Results show a speed‑up of 12–16× over the single‑core baseline, with the selective‑memory version achieving up to 70 % reduction in I/O time and a 30 % decrease in peak memory usage relative to the non‑aggregated parallel run. A second benchmark scales the reaction network to several thousand species and reactions, confirming that the pipeline maintains >90 % CPU utilization and that performance scales almost linearly with core count until memory bandwidth becomes limiting.
In the discussion, the authors attribute the observed gains to three factors: (i) FastFlow’s lock‑free communication eliminates costly mutex contention; (ii) selective memory’s data reduction minimizes cache pressure and disk writes; and (iii) the static farm pattern balances load evenly across cores when the number of trajectories vastly exceeds the number of workers. They acknowledge limitations: the current pipeline is static, so dynamic load‑balancing or adaptive worker counts are not supported; only basic statistics (mean, variance) are aggregated out‑of‑the‑box, requiring custom extensions for histograms, joint distributions, or time‑correlated metrics; and the approach assumes that the SSA workload dominates compute time, which may not hold for hybrid deterministic‑stochastic models.
Future work outlined includes integrating GPU‑accelerated SSA kernels, extending selective memory to support arbitrary user‑defined reducers, and exploring elastic scaling in cloud environments where cores can be provisioned on demand. The authors also propose coupling StochKit‑FF with parameter‑sweep frameworks to enable automated sensitivity analysis at unprecedented speed.
In conclusion, StochKit‑FF demonstrates that a carefully engineered parallel pipeline, combined with runtime selective aggregation, can transform the cost profile of exact stochastic simulations in systems biology. By delivering near‑linear speed‑up on commodity multicore hardware while drastically reducing memory and I/O demands, the toolkit makes large‑scale ensemble simulations—essential for robust statistical inference—practically feasible for researchers without access to high‑performance clusters.
Comments & Academic Discussion
Loading comments...
Leave a Comment