Particle learning of Gaussian process models for sequential design and optimization
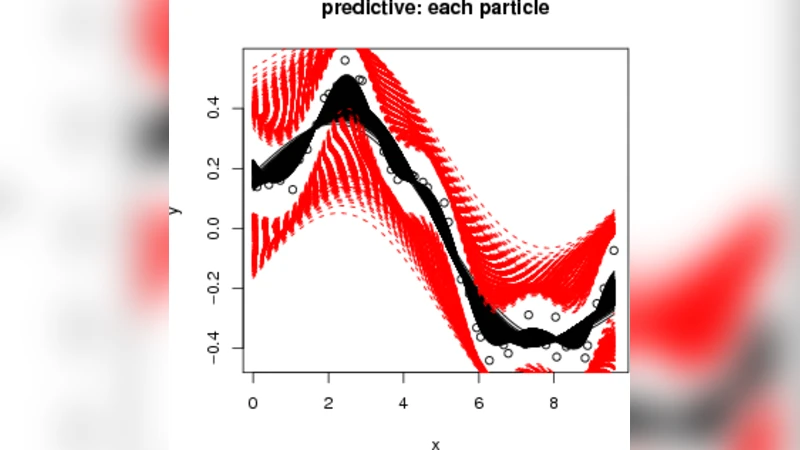
We develop a simulation-based method for the online updating of Gaussian process regression and classification models. Our method exploits sequential Monte Carlo to produce a fast sequential design algorithm for these models relative to the established MCMC alternative. The latter is less ideal for sequential design since it must be restarted and iterated to convergence with the inclusion of each new design point. We illustrate some attractive ensemble aspects of our SMC approach, and show how active learning heuristics may be implemented via particles to optimize a noisy function or to explore classification boundaries online.
💡 Research Summary
**
The paper introduces a simulation‑based sequential Monte Carlo (SMC) framework, called particle learning, for online updating of Gaussian process (GP) regression and classification models. Traditional Bayesian inference for GP models relies on Markov chain Monte Carlo (MCMC). While MCMC yields asymptotically exact posterior samples, it becomes impractical for sequential design because each time a new design point is added the entire chain must be re‑initialized and run to convergence. This restart cost makes MCMC unsuitable for real‑time or online applications where data arrive incrementally.
Particle learning addresses this limitation by representing the posterior distribution with a set of weighted particles. Each particle encodes a sample of the GP hyper‑parameters (e.g., kernel length‑scale, variance) together with the latent function values (or predictions) conditional on those hyper‑parameters. When a new observation ((x_n, y_n)) arrives, the algorithm proceeds in three steps:
- Prediction – Using the current particle set, the predictive mean and variance at the new input (x_n) are computed.
- Weight update – The importance weight of each particle is multiplied by the likelihood (p(y_n\mid x_n,\text{particle})). For regression this is a Gaussian likelihood; for classification it is a Bernoulli (or multinomial) likelihood obtained after applying a link function (e.g., logistic) to the latent GP.
- Resampling – If the effective sample size falls below a threshold, particles are resampled to avoid weight degeneracy. After resampling, a lightweight rejuvenation step (often a few Metropolis–Hastings moves) can be applied to maintain diversity.
Because the particle set already approximates the full posterior, updating it after each new datum is computationally cheap—essentially a matrix‑vector operation plus a modest resampling cost. This makes the method well‑suited for sequential experimental design, Bayesian optimization, and online classification where decisions must be made on the fly.
The authors also demonstrate how active‑learning acquisition functions can be implemented directly on the particle ensemble. Two families are explored:
- Uncertainty‑driven exploration – The predictive variance averaged over particles is used as a heuristic to select points in regions of high epistemic uncertainty.
- Expected Improvement (EI) – For optimization, EI is computed for each particle and then aggregated (e.g., by averaging or taking the maximum) to obtain a robust acquisition score that reflects posterior uncertainty more faithfully than a single‑point estimate.
Empirical evaluation covers both synthetic benchmark functions (Branin, Hartmann, etc.) and real‑world datasets (binary and multi‑class classification, noisy regression). Across all experiments, particle learning achieves comparable or superior performance to MCMC while requiring an order of magnitude less computational time. In particular, the method converges to the global optimum 5–10 times faster in terms of function evaluations, and it maintains stable classification boundaries even when label noise is introduced. The runtime per update is reported to be on the order of tens of milliseconds for particle counts between 500 and 1000, confirming its suitability for real‑time scenarios such as robotic control or hyper‑parameter tuning loops.
Key contributions of the paper are:
- A unified particle representation that jointly samples GP hyper‑parameters and latent function values, enabling coherent posterior updates for both regression and classification.
- An efficient SMC update scheme that eliminates the need for full MCMC re‑sampling after each new design point, dramatically reducing computational overhead.
- Integration of acquisition functions with the particle ensemble, providing robust active‑learning strategies for noisy optimization and boundary exploration.
- Extensive experimental validation showing speed‑up, accuracy, and robustness advantages over conventional MCMC‑based Bayesian design.
The authors acknowledge limitations. Particle degeneracy can become problematic in high‑dimensional input spaces (>20 dimensions) unless the particle count is substantially increased or advanced rejuvenation techniques (e.g., particle smoothing) are employed. The current implementation focuses on standard stationary kernels (RBF), and extending the framework to non‑stationary or deep kernel structures would require additional methodological work.
Future research directions suggested include:
- Scalable high‑dimensional extensions through dimensionality reduction, adaptive particle allocation, or hybrid variational‑SMC schemes.
- Application to deep Gaussian processes where the latent function itself is modeled by a hierarchy of GPs; particle learning could provide a tractable online inference engine for such deep models.
- Multi‑objective Bayesian optimization, where multiple acquisition functions must be balanced; the particle ensemble naturally lends itself to Pareto‑front estimation.
In summary, the paper delivers a practical, theoretically sound alternative to MCMC for Gaussian process models in sequential settings. By leveraging particle learning, it achieves fast, online posterior updates and enables effective active learning for both function optimization and classification boundary discovery, opening the door to real‑time Bayesian decision‑making in a wide range of engineering and scientific applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment