From statistical mechanics to information theory: understanding biophysical information-processing systems
These are notes for a set of 7 two-hour lectures given at the 2010 Summer School on Quantitative Evolutionary and Comparative Genomics at OIST, Okinawa, Japan. The emphasis is on understanding how biological systems process information. We take a physicist’s approach of looking for simple phenomenological descriptions that can address the questions of biological function without necessarily modeling all (mostly unknown) microscopic details; the example that is developed throughout the notes is transcriptional regulation in genetic regulatory networks. We present tools from information theory and statistical physics that can be used to analyze noisy nonlinear biological networks, and build generative and predictive models of regulatory processes.
💡 Research Summary
The manuscript is a set of lecture notes originally delivered at the 2010 Summer School on Quantitative Evolutionary and Comparative Genomics held at the Okinawa Institute of Science and Technology. Its primary aim is to show how physicists can approach the problem of biological information processing by constructing simple, phenomenological models that capture the essential functional behavior of complex systems without requiring a detailed microscopic description. The central example throughout the notes is transcriptional regulation in genetic regulatory networks, but the methods are presented as broadly applicable to any noisy, nonlinear biological circuit.
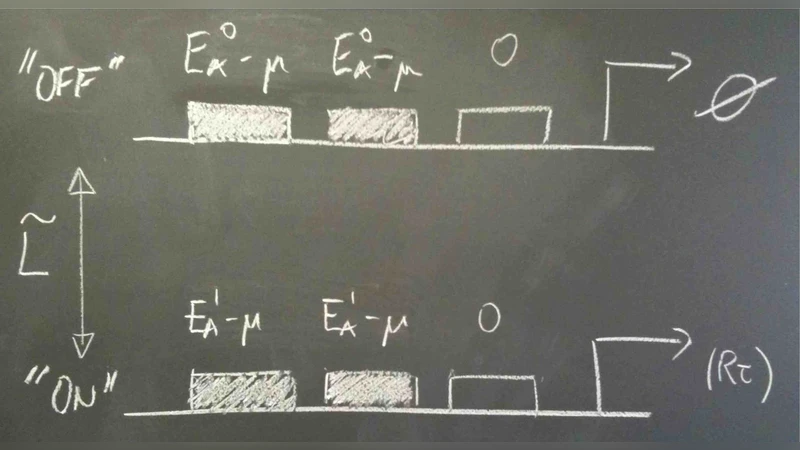
The first part introduces the conceptual framework: biological systems store and transmit information much like engineered communication channels, with DNA sequences acting as digital codes and transcription factors (TFs) serving as signal carriers. The authors emphasize that the binding of a TF to a promoter is a stochastic event governed by thermodynamic free‑energy differences, and that the resulting probability distribution can be described using Boltzmann statistics. By treating the TF‑DNA interaction as a statistical‑mechanical system, one can write down an explicit expression for the occupancy of a binding site as a function of TF concentration, temperature, and binding energy.
The second part brings in the tools of information theory. Entropy quantifies the uncertainty of the input (TF concentration) and output (gene expression level) distributions, while mutual information measures how much knowledge of the input reduces uncertainty about the output. The authors derive the channel capacity of a transcriptional regulatory element by optimizing the input distribution under realistic noise constraints. They show that biological systems often operate near this theoretical limit, suggesting evolutionary pressure to maximize information transmission.
In the third part the authors discuss how to fit these abstract models to real data. They adopt a Bayesian inference framework, assigning priors to parameters such as binding free energy, intrinsic noise amplitude, and cooperativity coefficients. Markov‑Chain Monte Carlo (MCMC) sampling yields posterior distributions that capture parameter uncertainty. Cross‑validation and bootstrapping are used to guard against over‑fitting and to assess predictive performance on independent datasets. The notes illustrate the workflow with published TF‑dose‑response curves, demonstrating that the fitted models can reproduce observed input‑output relationships and predict the effects of mutations or changes in cellular context.
Finally, the manuscript explores extensions beyond transcriptional regulation. The same statistical‑mechanical and information‑theoretic formalism can be applied to neural spike‑train coding, immune receptor–antigen recognition, and signaling cascades involving post‑translational modifications. By focusing on the macroscopic quantities that are experimentally accessible—such as mean expression levels, variances, and correlation functions—the authors argue that one can build generative, predictive models that are both tractable and biologically insightful.
Overall, the work showcases a powerful interdisciplinary toolkit: statistical mechanics provides a principled way to model stochastic binding events, while information theory supplies quantitative metrics for assessing the fidelity of biological communication. Together they enable researchers to move from descriptive biology toward a predictive, physics‑based understanding of how living systems process information in the presence of noise and limited resources.
Comments & Academic Discussion
Loading comments...
Leave a Comment