Revisiting Date and Party Hubs: Novel Approaches to Role Assignment in Protein Interaction Networks
The idea of ‘date’ and ‘party’ hubs has been influential in the study of protein-protein interaction networks. Date hubs display low co-expression with their partners, whilst party hubs have high co-expression. It was proposed that party hubs are local coordinators whereas date hubs are global connectors. Here we show that the reported importance of date hubs to network connectivity can in fact be attributed to a tiny subset of them. Crucially, these few, extremely central, hubs do not display particularly low expression correlation, undermining the idea of a link between this quantity and hub function. The date/party distinction was originally motivated by an approximately bimodal distribution of hub co-expression; we show that this feature is not always robust to methodological changes. Additionally, topological properties of hubs do not in general correlate with co-expression. Thus, we suggest that a date/party dichotomy is not meaningful and it might be more useful to conceive of roles for protein-protein interactions rather than individual proteins. We find significant correlations between interaction centrality and the functional similarity of the interacting proteins.
💡 Research Summary
The paper revisits the widely cited “date‑hub” versus “party‑hub” dichotomy that has shaped much of the thinking about protein‑protein interaction (PPI) networks. In the original formulation, hubs (high‑degree proteins) were split into two groups based on the correlation of their gene expression with that of their interaction partners. Party hubs were thought to be locally coordinated, showing high co‑expression with many partners, whereas date hubs were considered global connectors, displaying low co‑expression and linking disparate functional modules. The authors set out to test whether this binary classification truly reflects network topology, functional importance, or biological reality.
Using two well‑characterized interactomes (human and yeast), they first reproduced the hub‑co‑expression analysis. They found that the apparent bimodal distribution of hub‑partner expression correlation—used as the basis for the date/party split—is highly sensitive to methodological choices such as missing‑value imputation, log‑transformation, and normalization. When alternative preprocessing pipelines are applied, the bimodality disappears and the distribution becomes essentially unimodal. This suggests that the original dichotomy may be an artifact of data handling rather than a robust biological signal.
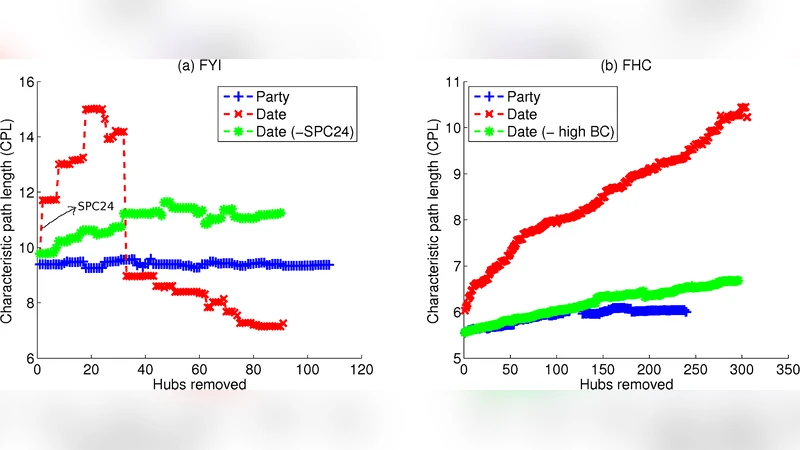
Next, the authors examined the contribution of hubs to overall network connectivity. By iteratively deleting individual hubs and measuring changes in average shortest‑path length and the size of the largest connected component, they discovered that only a very small fraction (≈5 % of all hubs) are truly critical for maintaining global connectivity. These “ultra‑central” hubs have exceptionally high degree and betweenness, and their removal fragments the network dramatically. Importantly, these ultra‑central hubs do not belong to the low‑co‑expression (date) group; in fact, they tend to have higher than average expression correlation with their partners. Consequently, the claim that date hubs are the primary global connectors is not supported when the analysis is restricted to the hubs that actually matter for network integrity.
The study also probed whether standard topological metrics (clustering coefficient, degree assortativity, eigenvector centrality, etc.) correlate with expression co‑variation. Across both organisms, no consistent or statistically significant relationships emerged. This further weakens the argument that a hub’s “date” or “party” status can be inferred from its position in the network topology.
To move beyond the protein‑centric view, the authors introduced an “interaction centrality” measure that evaluates the importance of each edge rather than each node. Interaction centrality quantifies how much the removal of a specific protein‑protein link would increase the overall network distance or disrupt connectivity. When they compared interaction centrality scores with functional similarity metrics (Gene Ontology term overlap, shared KEGG pathways), a strong positive correlation emerged: edges with high centrality tend to connect proteins that are functionally more similar. This finding suggests that the functional role of a PPI network is better captured by the properties of its interactions than by the attributes of individual hub proteins.
In summary, the paper presents four key conclusions: (1) the date/party hub classification is not robust; it depends on data preprocessing and does not survive methodological variation; (2) only a tiny subset of ultra‑central hubs drives network connectivity, and these hubs are not characterized by low expression correlation; (3) conventional topological descriptors of hubs do not predict co‑expression patterns; and (4) edge‑level centrality provides a more biologically meaningful metric, linking network structure to functional similarity. The authors therefore advocate a shift from a protein‑centric dichotomy to an interaction‑centric framework for interpreting PPI networks. This perspective has implications for network‑based disease gene prioritization, modular decomposition, and the design of experiments aimed at perturbing critical interactions rather than targeting individual “hub” proteins.
Comments & Academic Discussion
Loading comments...
Leave a Comment