Two-Domain DNA Strand Displacement
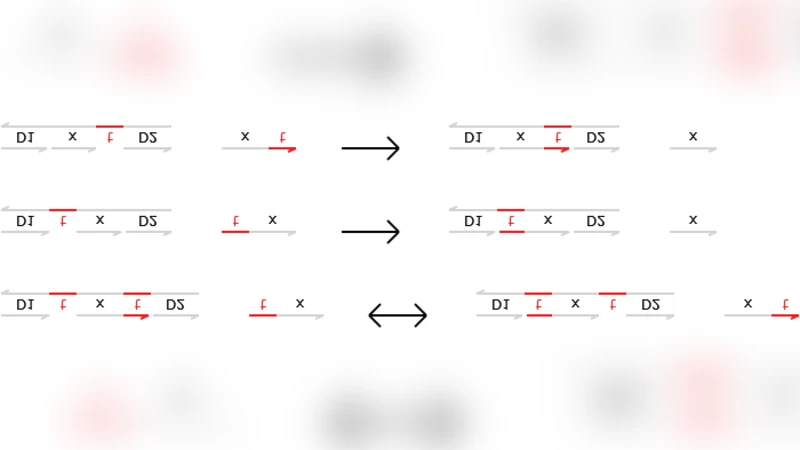
We investigate the computing power of a restricted class of DNA strand displacement structures: those that are made of double strands with nicks (interruptions) in the top strand. To preserve this structural invariant, we impose restrictions on the single strands they interact with: we consider only two-domain single strands consisting of one toehold domain and one recognition domain. We study fork and join signal-processing gates based on these structures, and we show that these systems are amenable to formalization and to mechanical verification.
💡 Research Summary
The paper investigates a highly constrained class of DNA strand‑displacement (DSD) systems in order to obtain a model that is both experimentally tractable and amenable to formal verification. The authors restrict the structural universe to “top‑nicked double‑stranded DNA” (ndsDNA), i.e., double‑stranded molecules in which only the top strand contains interruptions (nicks). Between any two nicks the top strand may contain at most two domains (a short toehold t and a long domain x, y, z, …). This restriction eliminates protruding single‑stranded overhangs, thereby reducing unintended interactions and simplifying the kinetic landscape.
In parallel, the signal molecules are limited to two‑domain single strands of the form t x or x t, where t is a single, universal toehold and x is a long domain that uniquely identifies the signal. The authors carefully analyze why other possible single‑strand patterns (e.g., x t y, t x t, or consecutive toeholds) would generate stable, non‑ndsDNA complexes and therefore must be excluded. The result is a minimal signal alphabet that guarantees that any interaction with an ndsDNA gate either proceeds through a well‑defined toehold‑mediated branch migration or remains reversible without forming spurious stable structures.
Within this framework the authors design four basic logical primitives:
-
Transducer T_xy – converts an input signal t x into an output signal t y. The construction uses a private domain a that links two nicked double strands; after two successive toehold exchanges the output t y is released while the auxiliary strands (including the private a‑containing strand) are re‑absorbed, leaving only inert dsDNA.
-
Fork F_xyz – splits a single input t x into two outputs t y and t z. It re‑uses the left half of the transducer and adds a symmetric right half, allowing simultaneous release of both outputs.
-
Catalyst C_xyz – consumes an input t x while preserving a catalyst signal t y, and produces a new output t z together with the unchanged catalyst t y. The right half of the fork is reused, and the catalyst strand is generated internally, eliminating the need for an external auxiliary strand.
-
Join J_xyz – merges two inputs t x and t y to produce a single output t z. A novel “garbage‑collector” sub‑structure (t†b y†t) is introduced to capture the co‑signal yt that would otherwise remain active after the reaction. This collector only activates after a private strand t b has been released, ensuring that the output is not prematurely scavenged.
All gates are built from nicked double strands that are concatenations of the elementary building blocks t, x, t x, x t, and x y. The authors prove that after any sequence of reactions the system returns to a configuration consisting solely of ndsDNA and inert single strands, i.e., no active toeholds or stray co‑signals persist. This “garbage‑free” property is crucial for scaling to larger networks because accumulated waste would otherwise generate reverse pressure on the desired forward reactions.
To reason formally about these systems the authors introduce Nick Algebra, a sub‑language of the broader DSD formalism. The syntax distinguishes single strands (S), double strands (D) possibly containing a nick (†), and multisets of strands (U). A domain‑isolation operator (ν x)U declares private domains that are guaranteed not to clash with any other part of the system, enabling compositional reasoning. The algebra includes a small set of reduction rules (D1–D4) that capture reversible toehold binding, irreversible branch migration, and the special nick‑mediated exchange steps. Because the language is deliberately minimal, it can be directly encoded into theorem provers such as Coq or Isabelle, allowing automated verification of properties like confluence, dead‑lock freedom, and the absence of residual active strands.
The discussion sections address both the practical and theoretical implications of the restrictions. Experimentally, ndsDNA structures are straightforward to synthesize; the single‑toehold design reduces the need for a large library of orthogonal toeholds, simplifying the design of large circuits. Theoretically, the authors show that despite the severe constraints, the constructed gates are computationally universal in the sense that they can simulate any Petri net (fork and join are sufficient). Moreover, the formal framework demonstrates that one can systematically extend the model to more complex structures if needed, but the current restricted set already covers a surprisingly rich class of computations.
In summary, the paper presents a disciplined approach to DNA‑based molecular computing: by limiting both the architecture of double‑stranded gates (top‑nicked) and the form of signal strands (two‑domain), the authors obtain a system that is experimentally realistic, computationally expressive, and formally verifiable. The combination of garbage‑free gate designs, a concise algebraic model, and mechanized proofs constitutes a significant step toward reliable, scalable molecular circuits.
Comments & Academic Discussion
Loading comments...
Leave a Comment