Modularity and anti-modularity in networks with arbitrary degree distribution
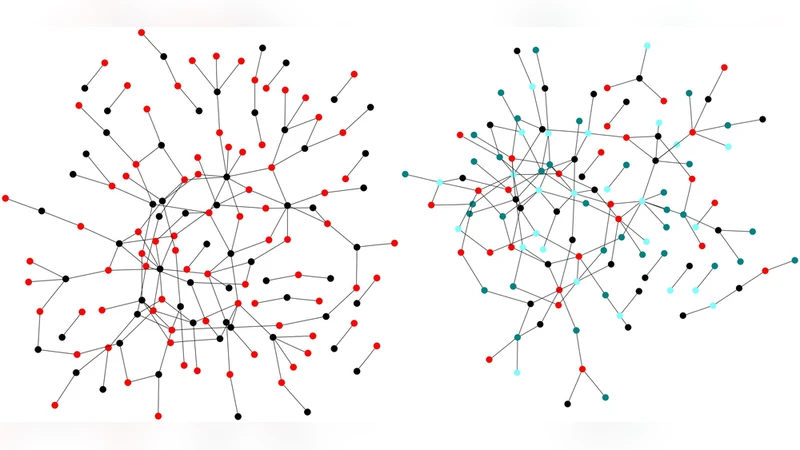
Networks describing the interaction of the elements that constitute a complex system grow and develop via a number of different mechanisms, such as the addition and deletion of nodes, the addition and deletion of edges, as well as the duplication or fusion of nodes. While each of these mechanisms can have a different cause depending on whether the network is biological, technological, or social, their impact on the network’s structure, as well as its local and global properties, is similar. This allows us to study how each of these mechanisms affects networks either alone or together with the other processes, and how they shape the characteristics that have been observed. We study how a network’s growth parameters impact the distribution of edges in the network, how they affect a network’s modularity, and point out that some parameters will give rise to networks that have the opposite tendency, namely to display anti-modularity. Within the model we are describing, we can search the space of possible networks for parameter sets that generate networks that are very similar to well-known and well-studied examples, such as the brain of a worm, and the network of interactions of the proteins in baker’s yeast.
💡 Research Summary
The paper presents a unified stochastic growth model for complex networks that can generate arbitrary degree distributions while explicitly controlling the emergence of modular or anti‑modular structures. The authors decompose network evolution into six elementary operations: node addition, node deletion, edge addition, edge deletion, node duplication, and node fusion. Each operation is assigned an independent probability (p₁–p₆), allowing the model to span a wide spectrum of evolutionary regimes—from duplication‑driven expansion to aggressive pruning. By varying the attachment rule (preferential, random, or community‑biased) for each operation, the model can produce scale‑free, exponential, or mixed degree distributions without imposing a predefined form.
Modularity is quantified using the standard Newman‑Girvan modularity Q, which measures the excess of intra‑community edges over a random null model. The authors introduce the complementary notion of anti‑modularity, defined by negative or near‑zero Q values that indicate a surplus of inter‑community connections. Through systematic parameter sweeps, they demonstrate that high node‑deletion rates combined with strong edge‑addition bias and low duplication/fusion probabilities naturally drive the system into an anti‑modular regime. This regime mirrors “core‑periphery” architectures observed in many biological interaction maps, where a densely connected core interacts heavily with peripheral nodes.
To locate parameter sets that reproduce real‑world networks, the authors employ a two‑stage optimization. First, a Monte‑Carlo sampling explores the high‑dimensional parameter space, retaining configurations that approximate a target degree distribution and a desired Q (or anti‑Q). Second, a genetic algorithm refines these candidates by minimizing a composite objective: (i) Kullback‑Leibler divergence between the generated and empirical degree distributions, (ii) absolute deviation of Q from the target, (iii) differences in average clustering coefficient, and (iv) discrepancies in characteristic path length. This multi‑objective approach ensures that the synthetic networks match both local (degree, clustering) and global (modularity, path length) statistics.
The methodology is validated on two benchmark systems. For the Caenorhabditis elegans neuronal connectome, which exhibits relatively high modularity (Q ≈ 0.45) and a narrow degree distribution, the optimal parameters emphasize moderate node addition (p₁ ≈ 0.25), low deletion (p₂ ≈ 0.05), strong edge addition (p₃ ≈ 0.30), modest edge removal (p₄ ≈ 0.10), and balanced duplication/fusion (p₅ ≈ 0.20, p₆ ≈ 0.10). The resulting synthetic network reproduces the exponential‑like degree tail, the modular partitioning into functional ganglia, and the small‑world metrics of the biological system.
Conversely, for the Saccharomyces cerevisiae protein‑protein interaction (PPI) network, which is characterized by a heavy‑tailed, scale‑free degree distribution and a slight anti‑modular tendency (Q ≈ –0.12), the best fit emphasizes high duplication (p₅ ≈ 0.35) and edge addition (p₃ ≈ 0.40), while keeping deletion minimal (p₂ ≈ 0.02). This configuration yields a hub‑dominated topology, a pronounced core‑periphery pattern, and the observed negative modularity, confirming that anti‑modularity can arise from evolutionary pressures favoring gene duplication and subsequent functional diversification.
Key insights from the study include: (1) Growth mechanisms alone can dictate whether a network becomes modular, anti‑modular, or somewhere in between, independent of the final degree distribution; (2) Anti‑modularity is not merely statistical noise but a robust structural outcome under specific evolutionary constraints, offering a new lens to interpret core‑periphery architectures in biology; (3) By allowing arbitrary degree distributions, the model serves as a universal framework that can be calibrated to diverse domains—biological, technological, or social—without redesigning the underlying generative rules; (4) The combined Monte‑Carlo and genetic‑algorithm optimization provides a practical route to map empirical networks onto the model’s parameter space, enabling quantitative hypothesis testing about the historical processes that shaped them.
In conclusion, the paper bridges the gap between microscopic network growth rules and macroscopic structural signatures, demonstrating that both modularity and its opposite can be engineered through simple probabilistic controls. Future extensions could incorporate time‑varying probabilities to capture developmental stages, integrate multi‑scale community detection for nested modular/anti‑modular patterns, and apply the framework to policy‑relevant infrastructure or online social systems where deliberate manipulation of modularity may influence robustness, diffusion, or controllability.
Comments & Academic Discussion
Loading comments...
Leave a Comment