Can entropy save bacteria?
This article presents a physical biology approach to understanding organization and segregation of bacterial chromosomes. The author uses a “piston” analogy for bacterial chromosomes in a cell, which leads to a phase diagram for the organization of two athermal chains confined in a closed geometry characterized by two length scales (length and width). When applied to rod-shaped bacteria such as Escherichia coli, this phase diagram predicts that, despite strong confinement, duplicated chromosomes will demix, i.e., there exists a primordial physical driving force for chromosome segregation. The author discusses segregation of duplicating chromosomes using the concentric-shell model, which predicts that newly synthesized DNA will be found in the periphery of the chromosome during replication. In contrast to chromosomes, these results suggest that most plasmids will be randomly distributed inside the cell because of their small sizes. An active partitioning system is therefore required for accurate segregation of low-copy number plasmids. Implications of these results are also sketched, e.g., on the role of proteins, segregation mechanisms for bacteria of diverse shapes, cell cycle of an artificial cell, and evolution.
💡 Research Summary
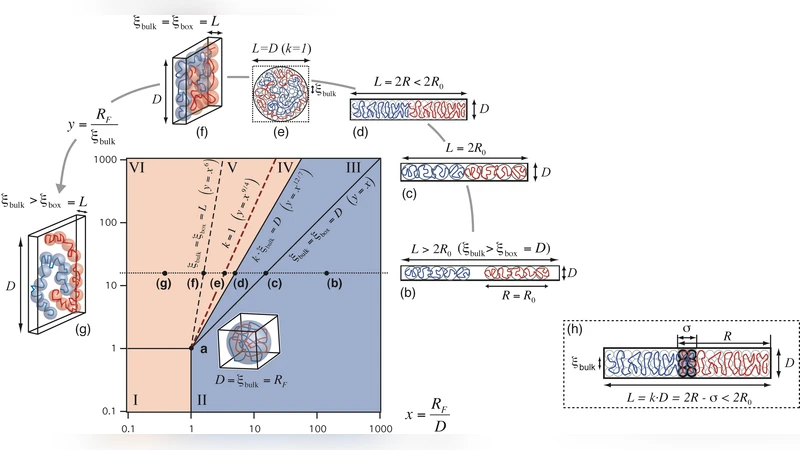
The paper presents a physical‑biology framework for understanding how bacterial chromosomes and plasmids become organized and segregated inside a confined cell. The author models each chromosome as an athermal polymer chain and treats the bacterial cell as a closed rectangular box characterized by two length scales: the cell’s longitudinal length (L_cell) and its transverse width (W_cell). By analogy to a piston, two such chains confined together experience an entropic pressure that drives them to minimize overlap. This leads to a dimensionless parameter (L/L_cell)·(R/W_cell), where L is the contour length of a chain and R reflects its effective thickness or flexibility. Plotting this parameter yields a phase diagram with two regimes: a “mix” region where chains interpenetrate and a “demix” region where they spontaneously separate.
When the model is applied to rod‑shaped bacteria such as Escherichia coli, the total length of a duplicated chromosome is comparable to the cell length, while the cell width is relatively small. Consequently, the dimensionless product falls squarely in the demix region, predicting that even in the absence of active mechanisms the two sister chromosomes will be pushed apart toward opposite cell poles. The author refines this picture with a concentric‑shell model: newly synthesized DNA is inserted into the outer shell of the existing chromosome, so that replication forks tend to reside near the periphery of the nucleoid. This prediction matches experimental observations of replication zones being close to the cell ends.
In contrast, low‑copy-number plasmids are much shorter than a chromosome. Their (L/L_cell)·(R/W_cell) values lie in the mix region, implying that pure entropy would leave them randomly distributed throughout the cytoplasm. Therefore, accurate segregation of such plasmids requires dedicated active partitioning systems (e.g., ParA/ParB) that bias their positioning.
The author extends the analysis to cells of different shapes (cylindrical, spherical, irregular). Increasing the cell width shifts the dimensionless parameter toward the mix region, suggesting that organisms with broader cells may rely more heavily on protein‑mediated segregation. Conversely, very narrow or elongated cells can exploit the entropic demixing force more effectively.
Implications for synthetic biology are also discussed. By designing artificial vesicles whose dimensions place chromosomal DNA in the demix regime, one could achieve passive chromosome segregation without complex motor proteins, offering a plausible route for early protocells to inherit genetic material reliably. The work thus bridges polymer physics, cell morphology, and evolutionary considerations, arguing that entropy provides a primordial driving force for chromosome segregation while highlighting why plasmids, due to their small size, must evolve active partitioning mechanisms.
Comments & Academic Discussion
Loading comments...
Leave a Comment