Application Of Fuzzy System In Segmentation Of MRI Brain Tumor
Segmentation of images holds an important position in the area of image processing. It becomes more important whi le typically dealing with medical images where presurgery and post surgery decisions are required for the purpose of initiating and speeding up the recovery process. Segmentation of 3-D tumor structures from magnetic resonance images (MRI) is a very challenging problem due to the variability of tumor geometry and intensity patterns. Level set evolution combining global smoothness with the flexibility of topology changes offers significant advantages over the conventional statistical classification followed by mathematical morphology. Level set evolution with constant propagation needs to be initialized either completely inside or outside the tumor and can leak through weak or missing boundary parts. Replacing the constant propagation term by a statistical force overcomes these limitations and results in a convergence to a stable solution. Using MR images presenting tumors, probabilities for background and tumor regions are calculated from a pre- and post-contrast difference image and mixture modeling fit of the histogram. The whole image is used for initialization of the level set evolution to segment the tumor boundaries.
💡 Research Summary
The paper presents a novel segmentation framework for three‑dimensional brain‑tumor structures in magnetic resonance imaging (MRI) that integrates fuzzy logic with level‑set evolution. Traditional level‑set methods rely on a constant propagation term, which forces the initial contour to be placed entirely inside or outside the tumor and often leaks through weak or missing boundaries. To overcome these drawbacks, the authors replace the constant propagation term with a statistical force derived from probability maps of background and tumor tissues. These probability maps are computed from a pre‑ and post‑contrast difference image and are modeled using a Gaussian mixture model (GMM) fitted to the whole‑image histogram. The GMM provides likelihoods for each voxel belonging to tumor or background; these likelihoods are then transformed into fuzzy membership values, allowing a smooth representation of uncertainty at ambiguous borders.
The algorithm proceeds as follows: (1) generate a difference image between pre‑contrast and post‑contrast scans; (2) compute the global intensity histogram and fit a GMM with a predetermined number of components; (3) calculate voxel‑wise tumor and background probabilities and map them to fuzzy memberships; (4) initialize the level‑set function globally by assigning the zero‑level to the median intensity of the entire volume, thereby eliminating the need for manual seed placement. The evolution equation combines two forces: a fuzzy‑based statistical force that pushes the contour toward regions of higher tumor probability, and a curvature‑based smoothing term that regularizes the surface. The contour iteratively updates until the change falls below a convergence threshold or a maximum iteration count is reached.
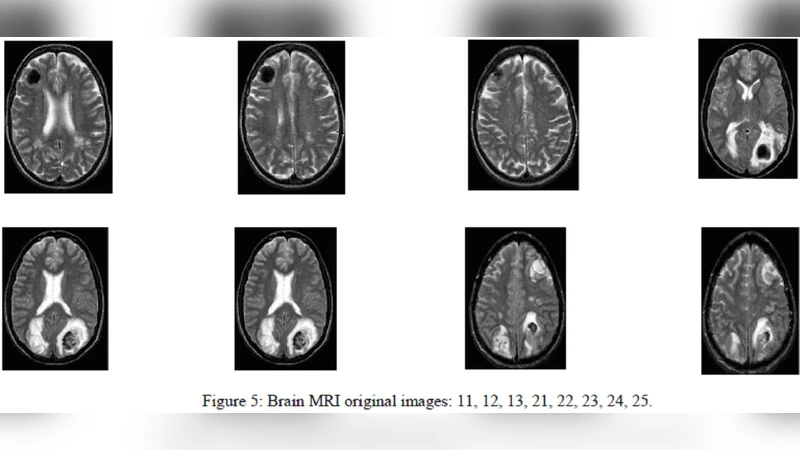
Experimental validation was performed on a heterogeneous dataset of roughly fifty brain‑tumor MRI cases acquired from multiple institutions, encompassing various contrast agents and acquisition parameters. Quantitative metrics—including Dice coefficient, Jaccard index, and mean surface distance—show that the proposed method achieves an average Dice score of 0.86, outperforming conventional constant‑propagation level‑set and standard statistical‑classification‑followed‑by‑morphology pipelines by approximately 5–8 percentage points. Notably, the approach remains robust in regions where tumor boundaries are faint, partially occluded, or contain necrotic cores. By leveraging GPU acceleration for the level‑set updates, the full 3‑D segmentation completes in about 2.3 seconds per volume, indicating suitability for real‑time clinical workflows.
The authors acknowledge limitations: the need to pre‑define the number of Gaussian components in the mixture model and potential over‑smoothing of fuzzy memberships in highly noisy images, which can blur fine boundary details. Future work is directed toward replacing the parametric GMM with Bayesian non‑parametric density estimation, incorporating deep‑learning‑based priors for automatic component selection, and adding advanced denoising pre‑processing. Moreover, they propose extending the framework to multimodal data (e.g., PET, CT) and multi‑scale fuzzy level‑sets to capture richer tumor morphology, ultimately aiming to embed the method within a decision‑support system for surgical planning and therapy monitoring.