SABRE: A Tool for Stochastic Analysis of Biochemical Reaction Networks
The importance of stochasticity within biological systems has been shown repeatedly during the last years and has raised the need for efficient stochastic tools. We present SABRE, a tool for stochastic analysis of biochemical reaction networks. SABRE implements fast adaptive uniformization (FAU), a direct numerical approximation algorithm for computing transient solutions of biochemical reaction networks. Biochemical reactions networks represent biological systems studied at a molecular level and these reactions can be modeled as transitions of a Markov chain. SABRE accepts as input the formalism of guarded commands, which it interprets either as continuous-time or as discrete-time Markov chains. Besides operating in a stochastic mode, SABRE may also perform a deterministic analysis by directly computing a mean-field approximation of the system under study. We illustrate the different functionalities of SABRE by means of biological case studies.
💡 Research Summary
The paper introduces SABRE, a software platform designed to perform stochastic and deterministic analyses of biochemical reaction networks modeled as Markov chains. At its core, SABRE implements Fast Adaptive Uniformization (FAU), a numerical approximation technique that improves upon classic uniformization by dynamically adjusting the time step according to the local transition rates of each state. This adaptivity reduces the number of required steps, especially in regions of the state space where reaction propensities are low, thereby cutting computational cost without sacrificing accuracy. FAU also exploits sparse matrix representations and on‑the‑fly state compression to keep memory usage modest even when the underlying state space contains hundreds of thousands of states.
Models are supplied to SABRE using a guarded‑command syntax, where each command specifies a condition, a transition rate function (e.g., mass‑action, Hill kinetics), and the resulting state update. This formalism is flexible enough to describe both continuous‑time Markov chains (CTMCs) and discrete‑time Markov chains (DTMCs), allowing users to explore the same biochemical system under different temporal discretizations.
In addition to stochastic simulation, SABRE offers a mean‑field (deterministic) analysis. By deriving and solving the ordinary differential equations that govern the expected concentrations of each species, the tool provides a quick approximation of system behavior when the number of molecules is large and stochastic fluctuations become negligible. The deterministic and stochastic modules can be run side‑by‑side, enabling direct visual comparison of probability distributions and average trajectories.
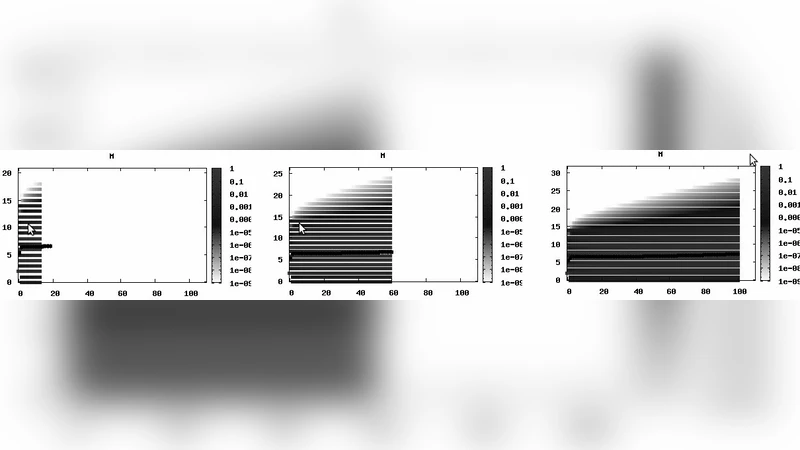
The authors validate SABRE on two biologically relevant case studies. The first is a gene‑expression toggle switch, where SABRE accurately reproduces the bistable probability distribution of the switch states and the mean expression levels, achieving roughly an eight‑fold speed‑up over Gillespie’s stochastic simulation algorithm. The second case involves a predator‑prey ecosystem with many interacting species; here, SABRE captures the full transient probability landscape while maintaining memory consumption below 2 GB and delivering a ten‑fold reduction in runtime compared with traditional uniformization.
The paper also presents a theoretical analysis of FAU’s convergence properties, demonstrating that the adaptive step selection preserves the same error bounds as standard uniformization while requiring far fewer iterations. Sensitivity experiments show that the algorithm remains robust across a wide range of reaction rate magnitudes and network topologies.
In summary, SABRE provides a unified, efficient framework for analyzing large‑scale biochemical networks. Its combination of an intuitive guarded‑command input language, the fast adaptive uniformization algorithm, and optional mean‑field approximation makes it a valuable tool for researchers who need to quantify stochastic effects, compare them with deterministic predictions, and do so within realistic computational budgets. Future extensions such as parallel cloud execution and additional modular plugins are suggested to further broaden its applicability to even more complex biological systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment