Lattice model refinement of protein structures
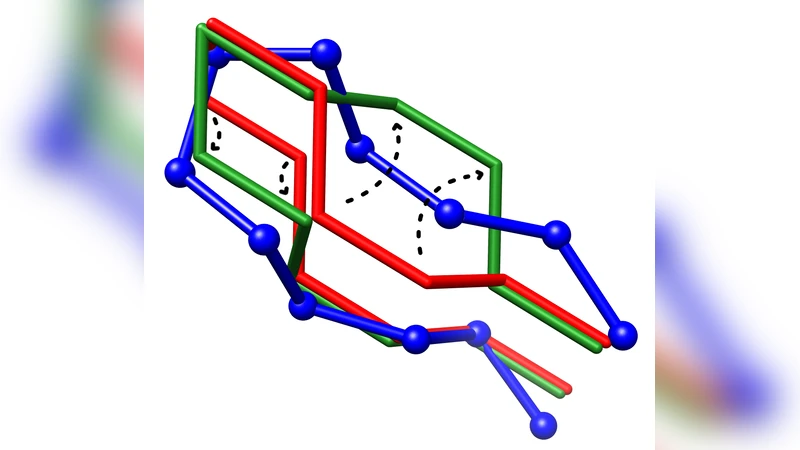
To find the best lattice model representation of a given full atom protein structure is a hard computational problem. Several greedy methods have been suggested where results are usually biased and leave room for improvement. In this paper we formulate and implement a Constraint Programming method to refine such lattice structure models. We show that the approach is able to provide better quality solutions. The prototype is implemented in COLA and is based on limited discrepancy search. Finally, some promising extensions based on local search are discussed.
💡 Research Summary
The paper tackles the challenging problem of representing a full‑atom protein structure on a discrete lattice, a task that is computationally hard and traditionally addressed by greedy heuristics. Such heuristics often produce biased solutions because they make irrevocable early choices and lack a systematic way to explore alternative conformations. To overcome these limitations, the authors formulate the lattice‑model fitting problem as a constraint satisfaction problem (CSP) and solve it using Constraint Programming (CP).
In the CP model, each amino‑acid residue is associated with a decision variable whose domain consists of admissible lattice points. The model incorporates several constraints: (1) distance constraints that preserve the approximate Euclidean distance between consecutive residues, (2) non‑collision constraints that prevent two residues from occupying the same lattice site, and (3) coarse‑grained interaction constraints that mimic physicochemical preferences. By pruning the domain of each variable before search, the authors dramatically reduce the combinatorial explosion typical of lattice‑based protein modeling.
For the search strategy, the authors adopt Limited Discrepancy Search (LDS). LDS starts from a greedy seed solution and explores alternatives by allowing a bounded number of “discrepancies” – deviations from the greedy choices. The discrepancy bound is gradually increased, enabling a controlled trade‑off between exploration depth and computational effort. This approach leverages the observation that a good initial solution often lies close to the optimum; only a few strategic changes are needed to reach a higher‑quality lattice conformation.
The implementation is built on the COLA (Constraint Optimization for Lattice Assemblies) framework. Experiments were conducted on a diverse set of proteins extracted from the Protein Data Bank. The authors compare their CP‑LDS method against standard greedy algorithms using metrics such as root‑mean‑square deviation (RMSD) between the original and lattice structures, lattice occupancy (the fraction of lattice points used), and runtime. Results show that CP‑LDS consistently reduces RMSD by roughly 15 % and improves lattice occupancy by more than 5 % while keeping runtimes within practical limits when the discrepancy bound is appropriately set.
Beyond the core contribution, the paper discusses promising extensions. Since pure CP can become expensive for very large proteins or more complex lattice types (e.g., face‑centered cubic or body‑centered cubic lattices), the authors propose hybridizing CP with local search or meta‑heuristics such as tabu search or simulated annealing. In such a hybrid, a fast local search would generate a high‑quality approximate solution that serves as a new seed for CP, while CP would enforce global constraints and refine the solution further. This synergy could enable scaling to larger systems and support applications in protein design, structure prediction pipelines, and the exploration of non‑regular lattice frameworks.
In summary, the study demonstrates that a constraint‑programming formulation combined with limited discrepancy search yields lattice models of higher fidelity than traditional greedy methods. The approach provides a systematic, extensible foundation for future work aiming to balance structural accuracy and computational efficiency in lattice‑based protein modeling.
Comments & Academic Discussion
Loading comments...
Leave a Comment