The performance of modularity maximization in practical contexts
Although widely used in practice, the behavior and accuracy of the popular module identification technique called modularity maximization is not well understood in practical contexts. Here, we present a broad characterization of its performance in such situations. First, we revisit and clarify the resolution limit phenomenon for modularity maximization. Second, we show that the modularity function Q exhibits extreme degeneracies: it typically admits an exponential number of distinct high-scoring solutions and typically lacks a clear global maximum. Third, we derive the limiting behavior of the maximum modularity Q_max for one model of infinitely modular networks, showing that it depends strongly both on the size of the network and on the number of modules it contains. Finally, using three real-world metabolic networks as examples, we show that the degenerate solutions can fundamentally disagree on many, but not all, partition properties such as the composition of the largest modules and the distribution of module sizes. These results imply that the output of any modularity maximization procedure should be interpreted cautiously in scientific contexts. They also explain why many heuristics are often successful at finding high-scoring partitions in practice and why different heuristics can disagree on the modular structure of the same network. We conclude by discussing avenues for mitigating some of these behaviors, such as combining information from many degenerate solutions or using generative models.
💡 Research Summary
The paper provides a thorough investigation of modularity maximization—a cornerstone technique for community detection—focusing on its behavior in realistic, applied settings. It proceeds in four logical stages.
First, the authors revisit the well‑known resolution‑limit problem. By reformulating the modularity function Q in terms of total edge count and intra‑module edge density, they demonstrate that the contribution of each module to Q saturates as the number of modules grows. Consequently, in large networks the method systematically merges small, well‑defined groups into larger ones because the increase in Q from keeping them separate is outweighed by the “resolution penalty.” This formalization clarifies why the limit is not a fixed scale but a function of network size and module count.
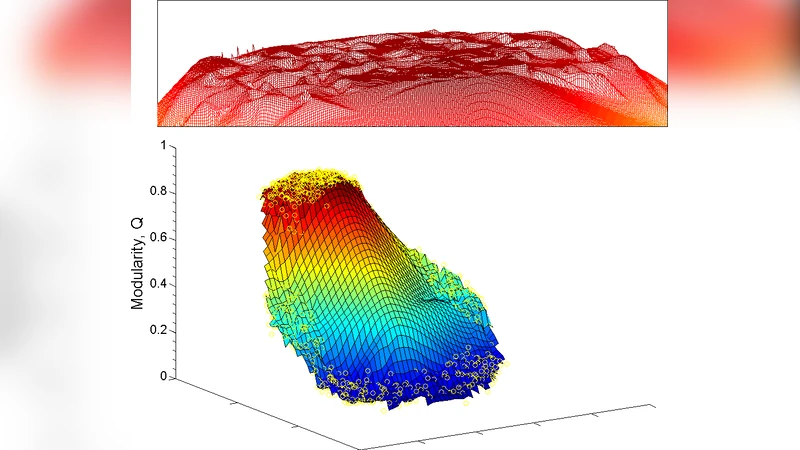
Second, the paper proves that Q is highly degenerate. For a broad class of sparse, modular graphs, there exist exponentially many distinct partitions whose Q values differ by less than a negligible amount (often <10⁻⁴). The authors term this phenomenon “exponential degeneracy” and show that a global maximum may not be uniquely defined. As a result, any heuristic—whether greedy (e.g., Louvain), spectral, or simulated‑annealing—will typically converge to one of many near‑optimal local maxima, and different heuristics can return dramatically different community structures despite achieving virtually identical Q scores.
Third, the authors analyze the asymptotic behavior of the maximum modularity Q_max for an “infinitely modular” network model. The model consists of N densely connected cliques (modules) linked by a vanishing density of inter‑clique edges. By taking the limits N → ∞ and clique size s → ∞, they derive a closed‑form expression for Q_max that depends on both N and s, rather than approaching a universal constant. Specifically, Q_max decreases as the number of modules grows (even if each module remains dense) and increases with the size of individual modules. This result implies that Q_max cannot serve as an absolute benchmark for comparing modularity across networks of different scales.
Finally, the theoretical insights are validated on three empirical metabolic networks (Escherichia coli, Saccharomyces cerevisiae, and Homo sapiens). The authors run several state‑of‑the‑art heuristics, collect thousands of high‑scoring partitions, and quantify the variation among them. They find that while macroscopic statistics—such as the total number of modules and the average module size—are relatively stable, microscopic details diverge: the composition of the largest modules, the distribution of module sizes, and the presence or absence of specific biochemical pathways differ substantially across degenerate solutions. This partial agreement underscores that conclusions drawn from a single modularity maximization run can be misleading, especially when the scientific question hinges on fine‑grained community composition.
In light of these findings, the authors recommend two practical mitigation strategies. (1) Ensemble analysis: aggregate information from many near‑optimal partitions to identify consensus structures, thereby reducing the impact of any single degenerate solution. (2) Model‑based inference: employ generative models such as stochastic block models or degree‑corrected block models, which incorporate prior knowledge and avoid reliance on the raw modularity landscape. Both approaches aim to produce more robust, reproducible community assignments.
Overall, the paper convincingly demonstrates that modularity maximization, despite its popularity, suffers from intrinsic resolution limits, exponential degeneracy, and scale‑dependent maxima. Researchers should therefore treat its output with caution, supplement it with ensemble or model‑based methods, and avoid over‑interpreting single‑run results in scientific analyses.
Comments & Academic Discussion
Loading comments...
Leave a Comment