Coupling of cytoplasm and adhesion dynamics determines cell polarization and locomotion
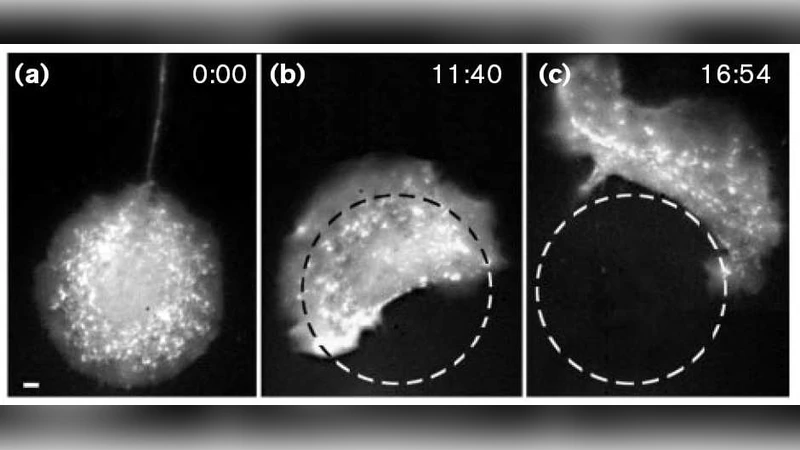
Observations of single epidermal cells on flat adhesive substrates have revealed two distinct morphological and functional states, namely a non-migrating symmetric unpolarized state and a migrating asymmetric polarized state. These states are characterized by different spatial distributions and dynamics of important biochemical cell components: F-actin and myosin-II form the contractile part of the cytoskeleton, and integrin receptors in the plasma membrane connect F-actin filaments to the substratum. In this way, focal adhesion complexes are assembled, which determine cytoskeletal force transduction and subsequent cell locomotion. So far, physical models have reduced this phenomenon either to gradients in regulatory control molecules or to different mechanics of the actin filament system in different regions of the cell. Here we offer an alternative and self-organizational model incorporating polymerization, pushing and sliding of filaments, as well as formation of adhesion sites and their force dependent kinetics. All these phenomena can be combined into a non-linearly coupled system of hyperbolic, parabolic and elliptic differential equations. Aim of this article is to show how relatively simple relations for the small-scale mechanics and kinetics of participating molecules may reproduce the emergent behavior of polarization and migration on the large-scale cell level.
💡 Research Summary
The paper addresses the fundamental question of how single epidermal cells on a flat adhesive substrate switch between a symmetric, non‑migratory state and an asymmetric, migratory state. While previous theoretical work has largely focused on gradients of regulatory molecules (e.g., Rho‑GTPases) or spatial variations in actin mechanics, the authors propose a self‑organizing model that explicitly couples three core physical processes: (1) actin filament polymerization and retrograde flow, (2) myosin‑II‑driven contractility, and (3) integrin‑mediated focal adhesion formation and force‑dependent turnover. Each process is described by a distinct class of partial differential equations—hyperbolic for filament transport, parabolic for contractile stress diffusion, and elliptic for the quasi‑static balance of adhesion forces. The equations are non‑linearly coupled through shared fields such as filament density, myosin concentration, and adhesion density, as well as through mechanical feedback: the force transmitted through adhesions modulates both polymerization rates and myosin activity, while the contractile stress influences adhesion reinforcement or disassembly.
The actin dynamics are modeled by a conservation law that includes a polymerization term proportional to the local G‑actin pool and a depolymerization term that depends on filament tension. Myosin‑II dynamics are captured by a diffusion‑reaction equation where contractile stress is proportional to myosin concentration and ATP hydrolysis rate, and the stress tensor feeds back into the actin transport equation. Focal adhesions are represented by a density field that obeys a force‑dependent kinetic law: the binding rate increases with the magnitude of traction force up to a saturation point, whereas the unbinding rate follows a Bell‑type exponential dependence on force. This captures the experimentally observed reinforcement of adhesions under load and rapid turnover when load is low.
Boundary conditions enforce force balance at the cell edge and no‑flux for actin and myosin at the membrane, while the substrate is treated as an elastic half‑space providing a linear relation between traction and substrate deformation. Numerical simulations are performed using a finite‑difference/finite‑element hybrid scheme on a two‑dimensional domain representing the cell footprint. Parameter values are drawn from literature (polymerization speed ≈0.1 µm s⁻¹, myosin stall force ≈1 pN, adhesion on‑rate and off‑rate constants calibrated to measured focal adhesion lifetimes).
The model reproduces two distinct steady‑states. In the symmetric state, actin polymerization, myosin contractility, and adhesion formation are spatially uniform, leading to a static cell. Small perturbations (e.g., stochastic fluctuations) destabilize this state, triggering a symmetry‑breaking bifurcation. The resulting polarized state exhibits a leading edge rich in polymerizing actin and nascent adhesions, while the rear accumulates myosin‑generated tension and mature, force‑reinforced adhesions. The rear adhesions gradually release, allowing the cell body to translocate forward. The speed and direction of migration are shown to depend sensitively on substrate stiffness: stiff substrates promote adhesion reinforcement, slowing migration, whereas compliant substrates favor rapid adhesion turnover and faster movement.
Comparison with experimental observations demonstrates that the model captures key hallmarks of cell migration: lamellipodial protrusion driven by actin polymerization, rear retraction mediated by myosin contractility, and the mechanosensitive maturation of focal adhesions. Perturbation experiments that inhibit integrin binding or myosin activity are recapitulated by setting the corresponding kinetic parameters to zero, which forces the system back into the non‑polarized state.
In conclusion, the authors provide a mathematically rigorous yet biologically grounded framework that integrates cytoskeletal mechanics and adhesion kinetics. By showing that relatively simple, experimentally measurable relations at the molecular level can give rise to emergent cell‑scale polarization and locomotion, the work bridges the gap between molecular biophysics and whole‑cell behavior. This approach opens avenues for predictive modeling of cell migration in development, wound healing, and cancer metastasis, and suggests that targeting the coupled dynamics of actin, myosin, and integrins could be an effective strategy for controlling cell motility.
Comments & Academic Discussion
Loading comments...
Leave a Comment