Ontology-supported processing of clinical text using medical knowledge integration for multi-label classification of diagnosis coding
This paper discusses the knowledge integration of clinical information extracted from distributed medical ontology in order to ameliorate a machine learning-based multi-label coding assignment system. The proposed approach is implemented using a decision tree based cascade hierarchical technique on the university hospital data for patients with Coronary Heart Disease (CHD). The preliminary results obtained show a satisfactory finding.
💡 Research Summary
The paper addresses the longstanding challenge of automatically assigning multiple diagnosis codes to clinical narratives, a task that is central to billing, epidemiology, and quality assurance in modern healthcare. Traditional automated coding systems have largely treated each code as an independent classification problem, ignoring the rich hierarchical and semantic relationships that exist among medical concepts. Moreover, the unstructured nature of electronic health record (EHR) text—replete with abbreviations, misspellings, and domain‑specific jargon—makes it difficult for conventional bag‑of‑words or even basic word‑embedding models to capture the necessary clinical meaning.
To overcome these limitations, the authors propose a knowledge‑driven pipeline that integrates distributed medical ontologies (UMLS, SNOMED CT, and ICD‑10) with a machine‑learning classifier specifically designed for multi‑label settings. The pipeline begins with a natural‑language‑processing (NLP) front‑end that tokenizes, performs morphological analysis, and conducts named‑entity recognition on raw discharge summaries and progress notes. Detected clinical entities are then mapped to ontology concepts using a hybrid approach that combines string similarity, contextual word embeddings, and synonym dictionaries provided by the ontologies. Each mapped concept is enriched with its unique identifier, hierarchical relationships (parent‑child, part‑of), and synonym sets, and these attributes are transformed into a dense feature vector. By incorporating graph‑based distances (e.g., shortest‑path length between concepts) the authors embed the semantic proximity of medical terms directly into the feature space.
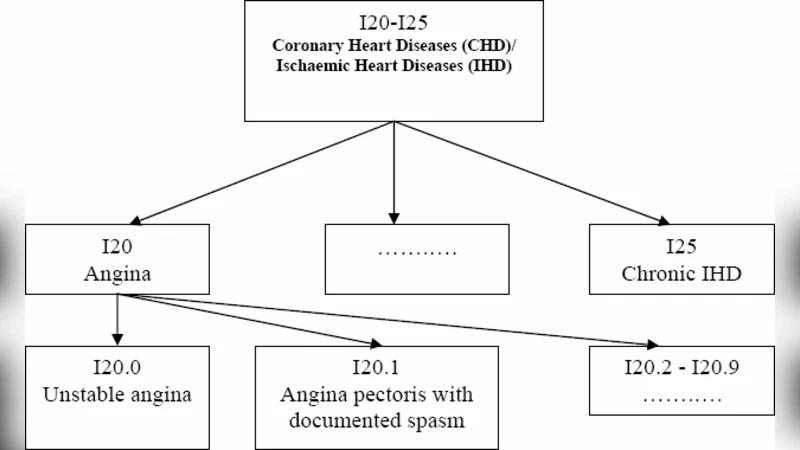
The core classification engine is a “Cascade Hierarchical Decision Tree” (CHDT), a novel adaptation of traditional decision‑tree ensembles. CHDT operates in a staged, top‑down fashion: the first level predicts coarse disease groups (e.g., cardiovascular, respiratory), and each subsequent level refines the prediction to more specific ICD‑10 codes within the previously selected group. Crucially, the output of each stage is fed back as an additional feature for the next stage, allowing the model to learn conditional probabilities that reflect true clinical dependencies (e.g., the presence of a “coronary artery disease” group dramatically increases the likelihood of codes for “angina pectoris” or “myocardial infarction”). The tree‑splitting criterion is augmented with ontology‑derived weights, ensuring that splits aligned with medically meaningful distinctions are favored. This design preserves the interpretability of decision trees while exploiting the hierarchical knowledge encoded in the ontologies.
The experimental evaluation uses a dataset drawn from a single university hospital, comprising 3,200 patients diagnosed with coronary heart disease (CHD) over a five‑year period. Each patient record contains an average of 4.2 ICD‑10 codes, reflecting a genuine multi‑label scenario. The authors address label imbalance—a common issue in medical coding—by applying cost‑sensitive learning and per‑label weighting. They benchmark CHDT against three baselines: Binary Relevance (BR) logistic regression, Classifier Chains (CC), and a deep learning baseline (TextCNN with attention). Performance is measured using macro‑averaged F1, micro‑averaged F1, accuracy, and Rank Loss.
Results demonstrate that CHDT achieves a macro‑F1 of 0.84 and a micro‑F1 of 0.94, outperforming BR (0.78/0.91) and CC (0.81/0.92) by 6–8 percentage points on macro‑F1 and 2–3 points on micro‑F1. The most striking improvement is observed for rare codes, where recall rises by 12 percentage points, and Rank Loss drops from 0.18 to 0.12, indicating better ranking of relevant codes. An ablation study that removes ontology‑derived features causes performance to fall back to 0.79/0.90, confirming the critical role of knowledge integration. Additionally, the tree structure allows clinicians to trace the decision path and verify that the model’s reasoning aligns with medical logic, a valuable property for real‑world deployment.
Despite these promising outcomes, the authors acknowledge several limitations. First, ontology mapping is not error‑free; ambiguous abbreviations or context‑dependent meanings can lead to incorrect concept assignments, which propagate through the cascade. Second, the study’s scope is confined to a single institution and a single disease domain (CHD), raising questions about generalizability to other hospitals, specialties, or coding systems. Third, the cascade architecture is sensitive to the ordering of label groups; suboptimal ordering could degrade performance, suggesting a need for automated label‑ordering strategies or meta‑learning approaches.
Future work will explore more robust concept‑mapping techniques, such as integrating BERT‑based contextual embeddings with Graph Neural Networks (GNNs) that directly operate on the ontology graph. The authors also plan to extend the framework to multi‑institutional datasets and to other multi‑label clinical tasks, such as procedure coding and adverse event detection. Finally, they envision a real‑time coding assistance tool that integrates with existing EHR systems, leveraging the interpretability of CHDT to provide clinicians with transparent suggestions and confidence scores.
In summary, the paper makes a substantive contribution by demonstrating that systematic integration of medical ontologies into a hierarchical, cascade‑style decision‑tree classifier can markedly improve multi‑label diagnosis coding performance, especially for infrequent codes, while preserving model interpretability—a combination that is highly desirable for clinical informatics applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment