A Formal Approach to Modeling the Memory of a Living Organism
We consider a living organism as an observer of the evolution of its environment recording sensory information about the state space X of the environment in real time. Sensory information is sampled and then processed on two levels. On the biological level, the organism serves as an evaluation mechanism of the subjective relevance of the incoming data to the observer: the observer assigns excitation values to events in X it could recognize using its sensory equipment. On the algorithmic level, sensory input is used for updating a database, the memory of the observer whose purpose is to serve as a geometric/combinatorial model of X, whose nodes are weighted by the excitation values produced by the evaluation mechanism. These values serve as a guidance system for deciding how the database should transform as observation data mounts. We define a searching problem for the proposed model and discuss the model’s flexibility and its computational efficiency, as well as the possibility of implementing it as a dynamic network of neuron-like units. We show how various easily observable properties of the human memory and thought process can be explained within the framework of this model. These include: reasoning (with efficiency bounds), errors, temporary and permanent loss of information. We are also able to define general learning problems in terms of the new model, such as the language acquisition problem.
💡 Research Summary
The paper proposes a formal, algorithmic model of memory for a living organism, treating memory as a dynamic, weighted graph (a database) that represents the organism’s perception of an external state space X. The organism O observes X in real time through a finite set of binary sensors. Each sensor partitions X into two complementary subsets; the collection of all such subsets is denoted H. H is closed under complementation, and the equivalence relation induced by H groups points of X that are indistinguishable to O into H‑visible states, forming a partition P(H).
The authors construct a graph Γ_H whose vertices are the elements of P(H). An edge connects two vertices P and Q precisely when there exists exactly one sensor h ∈ H such that P ⊆ h and Q ⊆ hᶜ (or vice‑versa). This construction is a special case of the dual graph of a space with walls, guaranteeing that Γ_H is connected and bipartite. The graph thus serves as a geometric/combinatorial model of the environment, while each vertex carries additional content: the empirical probabilities (or excitation values) that the organism has recorded for that H‑visible state.
Memory updates are formalized using category theory. Objects of the category C are weighted, connected bipartite graphs (the possible memory structures). Morphisms are surjective graph homomorphisms that may contract edges, representing the effect of adding new sensors (refining H to a larger ˜H) or merging states when evidence suggests they are indistinguishable. Thus, a refinement of the sensor set induces a morphism Γ_˜H → Γ_H, preserving the mapping from finer to coarser H‑visible states.
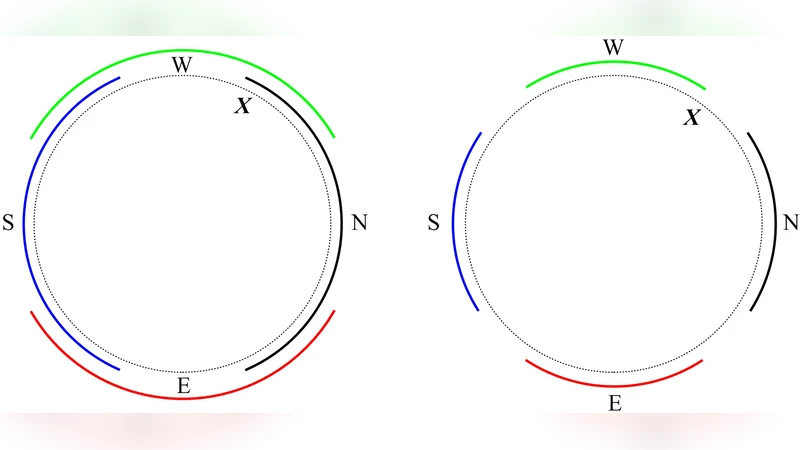
The paper examines computational aspects. As the number of sensors grows, Γ_H can become arbitrarily complex, potentially matching any large‑scale graph. Consequently, searching Γ_H (e.g., locating the current state, planning a path, or retrieving stored information) may become intractable. The authors illustrate this with a compass example: depending on the precision parameter ε, the graph can switch between a tree and a cycle, dramatically altering its topology for a tiny change in sensor accuracy. This illustrates how small sensory variations can force massive restructuring of the memory graph, raising questions about the feasibility of real‑time updates.
Beyond structural considerations, the model is used to explain several well‑known phenomena of human memory and cognition. Excitation values assigned by the biological evaluation mechanism act as weights guiding search and inference on Γ_H. Errors, temporary or permanent loss of information, and “glitches” are interpreted as mismatches between the organism’s recorded probabilities and the objective distribution, or as loss/rewiring of vertices and edges in the graph. The framework also accommodates learning problems: language acquisition, for instance, is modeled as the gradual addition of new sensors (grammatical cues) and the consequent refinement of the graph, together with weight adjustments reflecting exposure frequency.
Finally, the authors discuss implementation prospects. They suggest that a dynamic network of neuron‑like units could instantiate the graph, with units encoding vertices, synaptic connections encoding edges, and plasticity rules updating both topology and weights in response to sensory streams. Such a system would simultaneously capture the efficiency of biological memory, its susceptibility to errors, and its capacity for continual learning.
In summary, the paper introduces a mathematically rigorous, category‑theoretic description of memory as a weighted graph derived from binary sensory partitions, analyzes its computational properties, and demonstrates its explanatory power for a range of cognitive phenomena, while proposing a plausible neural implementation.
Comments & Academic Discussion
Loading comments...
Leave a Comment