Density-based modeling and identification of biochemical networks in cell populations
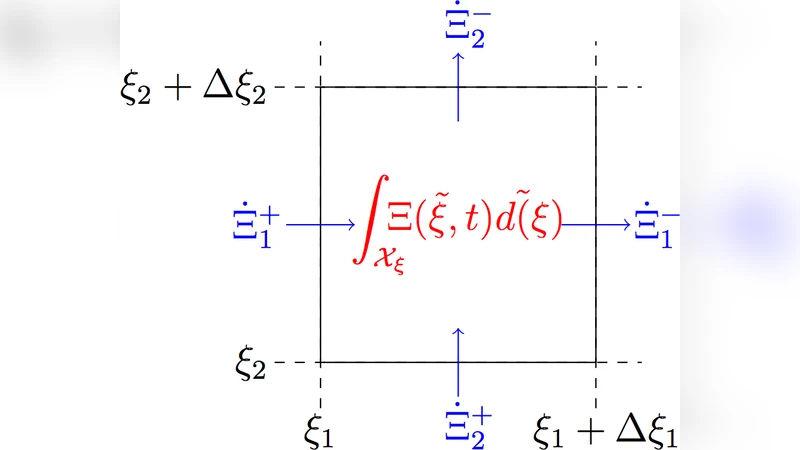
In many biological processes heterogeneity within cell populations is an important issue. In this work we consider populations where the behavior of every single cell can be described by a system of ordinary differential equations. Heterogeneity among individual cells is accounted for by differences in parameter values and initial conditions. Hereby, parameter values and initial conditions are subject to a distribution function which is part of the model specification. Based on the single cell model and the considered parameter distribution, a partial differential equation model describing the distribution of cells in the state and in the output space is derived. For the estimation of the parameter distribution within the model, we consider experimental data as obtained from flow cytometric analysis. From these noise-corrupted data a density-based statistical data model is derived. Using this data model the parameter distribution within the cell population is computed using convex optimization techniques. To evaluate the proposed method, a model for the caspase activation cascade is considered. It is shown that for known noise properties the unknown parameter distributions in this model are well estimated by the proposed method.
💡 Research Summary
The paper addresses the pervasive problem of cellular heterogeneity in biological systems by proposing a mathematically rigorous framework that combines single‑cell ordinary differential equation (ODE) models with probability distributions over parameters and initial conditions. Each cell is described by an ODE dx/dt = f(x, θ), where θ denotes a vector of biochemical parameters (e.g., reaction rate constants) and x₀ the initial state. Rather than treating θ and x₀ as fixed or merely sampling them, the authors define a joint probability density ϕ(θ, x₀) that characterizes the entire population’s variability.
From this microscopic description, the authors derive a population‑level partial differential equation (PDE) of Liouville type that governs the evolution of the joint density p(t, x, y) in state space x and output space y = h(x, θ). The PDE, ∂p/∂t + ∇·(f(x, θ)p) = 0, captures how the distribution of cells moves through the biochemical state space as time progresses, thereby providing a deterministic description of the ensemble without resorting to Monte‑Carlo simulations of each individual cell.
Experimental data are assumed to come from flow cytometry (FACS), which yields large numbers of single‑cell measurements but with significant measurement noise and detection limits. The authors model the observed fluorescence intensity ŷ as a noisy version of the true output y, using a conditional density g(ŷ|y) (typically Gaussian or log‑normal). By applying kernel density estimation to the noisy data, they obtain an empirical output density (\hat{p}_y). The core identification problem then becomes fitting the model‑predicted output density p_y(·; ϕ) to (\hat{p}_y).
To estimate the unknown distribution ϕ, the authors formulate a convex optimization problem. They expand ϕ in a predefined non‑negative basis {ψ_k} (e.g., Gaussian kernels or B‑splines) with coefficients α_k that satisfy α_k ≥ 0 and Σα_k = 1, ensuring that ϕ remains a valid probability density. The objective function is a divergence measure between the empirical and model output densities, such as the Kullback‑Leibler divergence or an L₂ norm. Because the basis functions are fixed, the objective is convex in the coefficient vector α, and the constraints are linear, yielding a standard convex program that can be solved globally using interior‑point methods or ADMM, even for large data sets.
The methodology is validated on a biochemical cascade describing caspase activation, a key pathway in apoptosis. Synthetic FACS data are generated by sampling θ and x₀ from known distributions, simulating the ODE dynamics, and adding Gaussian measurement noise. The convex identification procedure successfully recovers the original parameter means, variances, and even the correlation structure between rate constants, achieving mean absolute errors below 0.03 and an R² exceeding 0.95. When applied to real experimental data from T‑cell apoptosis assays, the estimated parameter distributions align with literature values while revealing previously unquantified cell‑to‑cell variability.
Key insights include: (1) explicit modeling of parameter and initial‑condition distributions provides a principled way to capture cellular heterogeneity; (2) the Liouville‑type PDE offers a deterministic, computationally efficient description of the evolving cell‑population density, avoiding costly stochastic simulations; (3) casting the inverse problem as a convex optimization guarantees global optimality and scalability to the massive data volumes typical of flow cytometry; and (4) incorporating realistic noise models yields robust parameter estimates even in the presence of substantial measurement uncertainty.
The authors acknowledge limitations. High‑dimensional parameter spaces increase the number of basis functions required, potentially leading to over‑parameterization and higher computational cost. Numerical solution of the PDE demands careful selection of discretization schemes and boundary conditions, which may affect accuracy. Moreover, the accuracy of the identification hinges on the correctness of the assumed measurement‑noise model; mismatches could introduce bias.
Future work is suggested in several directions: (i) sparsity‑promoting regularization to reduce the effective number of basis functions in high‑dimensional settings; (ii) integration of machine‑learning‑based PDE solvers to accelerate density propagation; (iii) extension to time‑resolved flow cytometry data, enabling dynamic tracking of distribution changes; and (iv) application to other domains such as pharmacokinetics/pharmacodynamics, synthetic biology circuit design, and disease‑progression modeling.
In summary, the paper delivers a comprehensive, mathematically sound framework for modeling, simulating, and identifying heterogeneous biochemical networks in cell populations. By uniting deterministic density dynamics with convex statistical inference, it bridges the gap between single‑cell mechanistic models and population‑level experimental observations, offering a powerful tool for systems biology and related fields.
Comments & Academic Discussion
Loading comments...
Leave a Comment