Modelling and Analysis of Biochemical Signalling Pathway Cross-talk
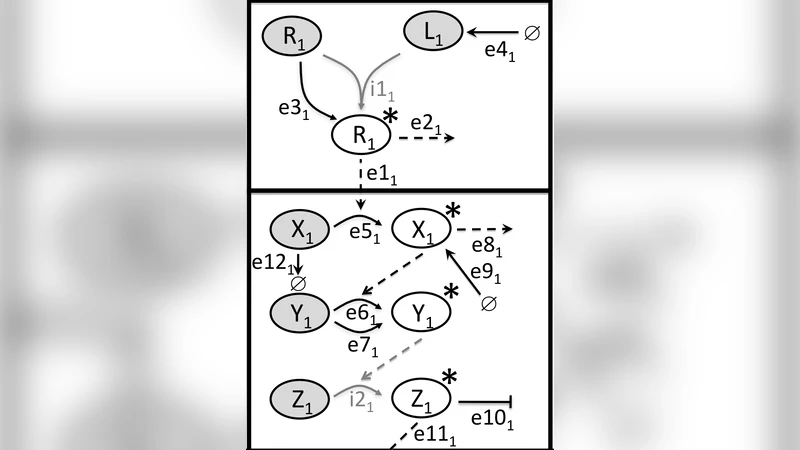
Signalling pathways are abstractions that help life scientists structure the coordination of cellular activity. Cross-talk between pathways accounts for many of the complex behaviours exhibited by signalling pathways and is often critical in producing the correct signal-response relationship. Formal models of signalling pathways and cross-talk in particular can aid understanding and drive experimentation. We define an approach to modelling based on the concept that a pathway is the (synchronising) parallel composition of instances of generic modules (with internal and external labels). Pathways are then composed by (synchronising) parallel composition and renaming; different types of cross-talk result from different combinations of synchronisation and renaming. We define a number of generic modules in PRISM and five types of cross-talk: signal flow, substrate availability, receptor function, gene expression and intracellular communication. We show that Continuous Stochastic Logic properties can both detect and distinguish the types of cross-talk. The approach is illustrated with small examples and an analysis of the cross-talk between the TGF-b/BMP, WNT and MAPK pathways.
💡 Research Summary
The paper presents a formal, modular approach to modeling biochemical signalling pathways and, in particular, the cross‑talk that occurs between them. The authors start from the premise that a signalling pathway can be represented as the synchronising parallel composition of a set of generic modules, each described by internal and external labels. Using the probabilistic model‑checking language PRISM, they encode these modules as continuous‑time Markov chains (CTMCs) with transition rates that capture kinetic parameters and stochastic fluctuations.
Cross‑talk is then defined as a combination of two algebraic operations on the modules: synchronisation, which forces two modules to fire the same labelled transition simultaneously, and renaming, which maps a label in one module to a label in another. By varying which labels are synchronised and which are renamed, the authors are able to capture five biologically relevant types of interaction:
- Signal‑flow cross‑talk – direct transmission of an output signal from one pathway into the input of another.
- Substrate‑availability cross‑talk – competition for a shared molecular substrate or intermediate.
- Receptor‑function cross‑talk – modulation of a receptor in one pathway by a ligand or effector from another.
- Gene‑expression cross‑talk – a transcription factor produced by one pathway regulating gene expression in a second pathway.
- Intracellular‑communication cross‑talk – indirect signalling mediated through organelles or spatial compartments, modelled by chains of synchronisations and renamings.
To detect and distinguish these patterns, the authors formulate a suite of Continuous Stochastic Logic (CSL) properties. For example, a property of the form `P≤0.1
Comments & Academic Discussion
Loading comments...
Leave a Comment