Conversion of graded to binary response in an activator-repressor system
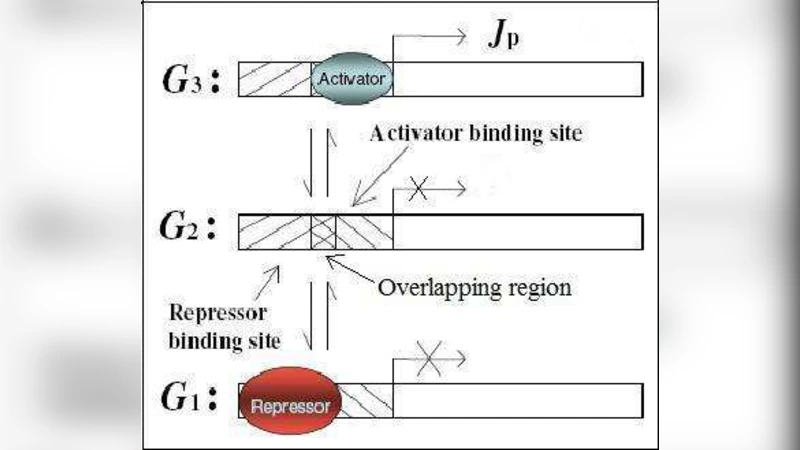
Appropriate regulation of gene expression is essential to ensure that protein synthesis occurs in a selective manner. The control of transcription is the most dominant type of regulation mediated by a complex of molecules such as transcription factors. In general, regulatory molecules are of two types: activator and repressor. Activators promote the initiation of transcription whereas repressors inhibit transcription. In many cases, they regulate the gene transcription on binding the promoter mutually exclusively and the observed gene expression response is either graded or binary. In experiments, the gene expression response is quantified by the amount of proteins produced on varying the concentration of an external inducer molecules in the cell. In this paper, we study a gene regulatory network where activators and repressors both bind the same promoter mutually exclusively. The network is modeled by assuming that the gene can be in three possible states: repressed, unregulated and active. An exact analytical expression for the steady-state probability distribution of protein levels is then derived. The exact result helps to explain the experimental observations that in the presence of activator molecules the response is graded at all inducer levels whereas in the presence of both activator and repressor molecules, the response is graded at low and high inducer levels and binary at an intermediate inducer level.
💡 Research Summary
The paper investigates how a gene regulatory system that uses a single promoter, to which an activator and a repressor bind mutually exclusively, can produce either graded (continuous) or binary (all‑or‑none) responses to an external inducer. The authors begin by framing the biological problem: transcriptional regulation is the primary means by which cells control protein synthesis, and the two canonical types of transcription factors—activators that promote initiation and repressors that block it—often compete for the same DNA site. Experimental observations have shown that when only activators are present the output (protein level) varies smoothly with inducer concentration, whereas the coexistence of activators and repressors can generate a switch‑like behavior at intermediate inducer levels while remaining graded at low and high concentrations.
To capture this phenomenon, the authors construct a minimal stochastic model in which the gene can occupy three discrete promoter states: repressed (R), unregulated (U), and active (A). In state R the transcription rate is zero, in U it is a basal rate α, and in A it is a maximal rate β (β > α). Binding of the activator and repressor to the promoter is assumed to follow Hill‑type kinetics with Hill coefficients n_A and n_R and dissociation constants K_A and K_R, respectively. The probability that the promoter is occupied by the activator at inducer concentration I is P_A(I)=I^{n_A}/(K_A^{n_A}+I^{n_A}), and similarly for the repressor P_R(I). Because the two factors are mutually exclusive, the transition rates between the three promoter states are functions of P_A and P_R, together with intrinsic on/off rates (k_on, k_off). Protein synthesis occurs only in states U and A, while degradation follows a first‑order rate γ.
The dynamics are described by a continuous‑time Markov master equation for the joint probability of promoter state and protein copy number. By introducing a probability‑generating function, the authors solve the steady‑state equation exactly. The solution reveals that the protein number distribution is a weighted sum of two Poisson distributions:
P(n) = w_R · Poisson(λ_R) + w_A · Poisson(λ_A)
where w_R and w_A are the steady‑state occupancies of the repressed and active states, and λ_R, λ_A are the corresponding mean protein numbers (λ_R = α/γ, λ_A = β/γ). The weight w_A depends non‑linearly on the inducer concentration through P_A and P_R.
When only the activator is present (P_R≈0), w_A increases monotonically with I, producing a single‑peak distribution that shifts smoothly—this is the graded response. When both factors are present, the competition creates a region of inducer concentration where w_R≈w_A≈0.5. In this region the two Poisson components have comparable weight, leading to a bimodal distribution: one peak near the low‑expression basin (repressed) and another near the high‑expression basin (active). This bimodality corresponds to the experimentally observed binary response. At very low or very high I, one component dominates, restoring a graded, unimodal profile.
Parameter sensitivity analysis shows that the width and position of the binary region are governed primarily by the ratio K_A/K_R and the Hill coefficients. A stronger repressor (smaller K_R) widens the binary window, while higher cooperativity (larger n_A or n_R) sharpens the transition. The ratio β/γ controls the separation between the two peaks: larger β or smaller γ yields more distinct high‑expression states.
To validate the analytical results, stochastic simulations using the Gillespie algorithm were performed across a range of inducer concentrations. The simulated protein histograms match the theoretical Poisson mixture, confirming that the exact solution captures the full stochastic behavior despite the simplifying assumptions of Markovian dynamics and a single promoter.
In the discussion, the authors emphasize that mutual exclusivity of activator and repressor binding is sufficient to generate a switch‑like response without invoking additional network motifs such as positive feedback. This insight has practical implications for synthetic biology: by tuning binding affinities and cooperativities, designers can engineer circuits that behave either as analog sensors (graded) or digital switches (binary) using the same genetic parts. The paper also outlines future extensions, including multiple promoters, cooperative binding between factors, and feedback loops, which could produce richer dynamical regimes such as oscillations or multistability.
Overall, the study provides a rigorous, analytically tractable framework that links molecular binding parameters to population‑level expression phenotypes, thereby deepening our understanding of how simple competitive interactions at a promoter can dictate whether a gene responds continuously or in an all‑or‑none fashion.
Comments & Academic Discussion
Loading comments...
Leave a Comment