Spectral Analysis of Virus Spreading in Random Geometric Networks
In this paper, we study the dynamics of a viral spreading process in random geometric graphs (RGG). The spreading of the viral process we consider in this paper is closely related with the eigenvalues of the adjacency matrix of the graph. We deduce new explicit expressions for all the moments of the eigenvalue distribution of the adjacency matrix as a function of the spatial density of nodes and the radius of connection. We apply these expressions to study the behavior of the viral infection in an RGG. Based on our results, we deduce an analytical condition that can be used to design RGG’s in order to tame an initial viral infection. Numerical simulations are in accordance with our analytical predictions.
💡 Research Summary
The paper investigates viral spreading dynamics on random geometric graphs (RGGs) by linking the process to the spectral properties of the adjacency matrix. Nodes are placed uniformly in a d‑dimensional Euclidean domain (the authors focus on the two‑dimensional unit square) and an undirected edge is created whenever the Euclidean distance between two nodes does not exceed a connection radius r. The average degree of such a graph is ρπr², where ρ=n/Area denotes the spatial node density.
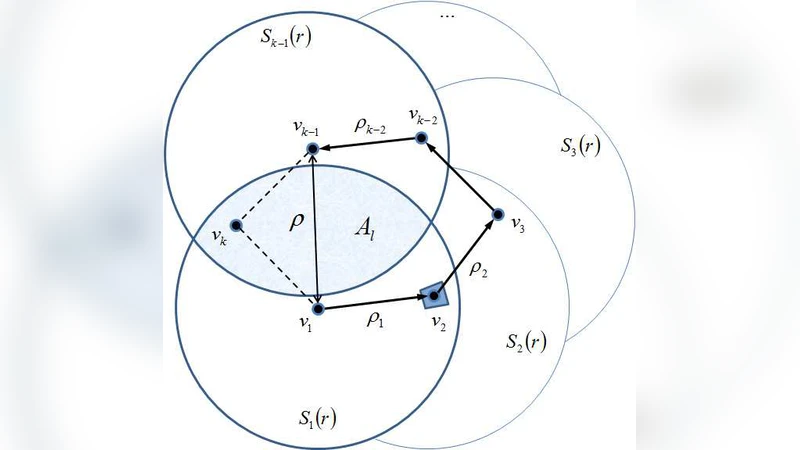
The core technical contribution is an exact derivation of all moments of the eigenvalue distribution μ(λ) of the adjacency matrix A. The k‑th moment Mₖ = ∫λᵏ dμ(λ) is interpreted as the expected number of closed walks of length k that start and end at the same vertex. By expressing each step of a walk as a geometric constraint (distance ≤ r) and integrating over the positions of the involved vertices, the authors obtain a closed‑form expression:
Mₖ = Cₖ (ρπr²)ᵏ,
where Cₖ depends only on the walk length k and the ambient dimension d. Cₖ can be written in terms of beta and gamma functions, and a simple recursion Cₖ₊₁ = Cₖ·(k+1)/(d·k) is provided, enabling efficient computation of high‑order moments.
Using the moment generating function M(z)=∑ₖMₖzᵏ, the authors bound the spectral radius λ_max (the largest eigenvalue of A). By applying Markov’s inequality together with Chebyshev polynomial approximations, they derive
λ_max ≤ ρπr² + √(2 ρπr²) + o(√(ρπr²)).
Thus the spectral radius grows linearly with the average degree, with a sub‑linear correction of order √(average degree).
The viral process is modeled as a standard SIS (susceptible–infected–susceptible) epidemic with infection probability β per edge per time step and recovery probability δ per infected node per time step. Classical spectral epidemic theory states that an infection persists if β/δ > 1/λ_max. Substituting the bound for λ_max yields an explicit design condition:
β/δ > 1/(ρπr²) ·
Comments & Academic Discussion
Loading comments...
Leave a Comment