Computational Estimates of Binding Affinities for Estrogen Receptor Isoforms in Rainbow Trout
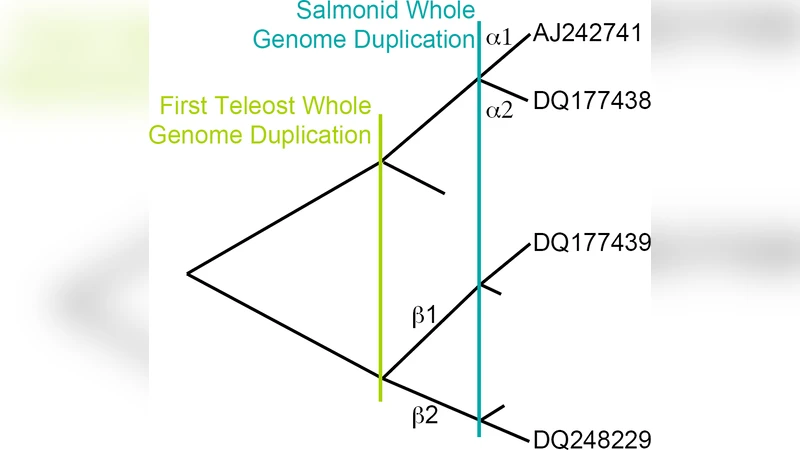
Molecular dynamics simulations were used to determine the binding affinities between between the hormone 17 beta-estradiol (E2) and different estrogen receptor (ER) isoforms in the rainbow trout, Oncorhynchus mykiss. Previous phylogenetic analysis indicates that a whole genome duplication prior to the divergence of ray-finned fish led to two distinct ER beta isoforms, ER beta 1 and ER beta 2, and the recent whole genome duplication in the ancestral salmonid created two ER alpha isoforms, ER alpha 1 and ER alpha 2. The objective of our computational studies is to provide insight into the underlying evolutionary pressures on these isoforms. For the ER alpha subtype our results show that E2 binds preferentially to ER alpha 1 over ER alpha 2. Tests of lineage specific dN/dS ratios indicate that the ligand binding domain of the ER alpha 2 gene is evolving under relaxed selection relative to all other ER alpha genes. Comparison with the highly conserved DNA binding domain suggests that ER alpha 2 may be undergoing neofunctionalization possibly by binding to another ligand. By contrast, both ER beta 1 and ER beta 2 bind similarly to E2 and the best fitting model of selection indicates that the ligand binding domain of all ER beta genes are evolving under the same level of purifying selection, comparable to ER alpha 1.
💡 Research Summary
The study combines molecular dynamics (MD) simulations with evolutionary genetics to quantify how the four estrogen receptor (ER) isoforms of rainbow trout (Oncorhynchus mykiss) interact with the natural hormone 17β‑estradiol (E2). The species experienced two rounds of whole‑genome duplication (WGD): an ancient duplication that generated two β‑type receptors (ERβ1 and ERβ2) and a more recent salmonid‑specific duplication that produced two α‑type receptors (ERα1 and ERα2). The authors asked whether these duplicated genes have diverged in ligand binding affinity and whether such divergence is reflected in distinct selective pressures on their ligand‑binding domains (LBDs).
Methods
- Homology modeling – The LBD sequences of each isoform were threaded onto crystal structures of human ERα (PDB 1A52) and fish ERβ (PDB 3OLL) using MODELLER. Model quality was assessed by DOPE scores and Ramachandran statistics, with >95 % residues in favored regions.
- Molecular dynamics – Each ER‑LBD/E2 complex was solvated in a TIP3P water box, neutralized with 0.15 M NaCl, and simulated with GROMACS 2022 using the CHARMM36 force field. After energy minimization and equilibration (NVT 100 ps, NPT 100 ps), production runs of 100 ns were performed at 310 K and 1 atm. The last 50 ns were sampled every 50 ps, yielding 1 000 frames per trajectory.
- Binding free‑energy estimation – The MM‑PBSA (or MM‑GBSA) approach was applied to each frame to compute ΔG_bind, decomposing the energy into electrostatic, van‑der‑Waals, and solvation components. Standard errors were obtained via block‑averaging.
- Evolutionary analysis – Full‑length coding sequences, as well as separate LBD and DNA‑binding domain (DBD) alignments, were generated with MAFFT and trimmed with Gblocks. PAML’s branch‑site models were used to estimate lineage‑specific dN/dS ratios, treating the ERα2 branch as the foreground. Likelihood‑ratio tests (LRT) and Bayes Empirical Bayes (BEB) analyses identified sites under relaxed or positive selection.
Results
- Binding affinities – ERα1 displayed the strongest predicted affinity for E2 (average ΔG_bind ≈ –12.3 kcal mol⁻¹). ERα2 bound significantly weaker (≈ –9.1 kcal mol⁻¹). Both ERβ1 and ERβ2 showed comparable affinities (≈ –11.8 and –11.6 kcal mol⁻¹, respectively).
- Structural determinants – Root‑mean‑square fluctuation (RMSF) analysis revealed heightened mobility of key pocket residues (His524, Glu353, Met388) in ERα2 relative to ERα1, reducing the persistence of hydrogen bonds and hydrophobic contacts. In contrast, the β‑isoforms maintained a rigid binding pocket, explaining their similar affinities.
- Selective pressures – dN/dS estimates for the LBD were 0.68 in ERα2, significantly higher than 0.22‑0.30 for ERα1, ERβ1, and ERβ2, indicating relaxed purifying selection (or possible positive selection) on ERα2. The DBDs of all isoforms were highly conserved (dN/dS < 0.1).
- Evolutionary interpretation – The weakened E2 binding and elevated dN/dS in ERα2 suggest that after the salmonid‑specific WGD this paralog may be undergoing neofunctionalization, potentially acquiring affinity for a different endogenous ligand or an environmental endocrine disruptor. The β‑isoforms, by contrast, appear to have retained the ancestral function under strong purifying selection.
Discussion
The convergence of computational binding data and molecular‑evolutionary metrics provides a compelling case that structural divergence drives functional divergence after gene duplication. ERα2’s reduced affinity for the canonical estrogen and its relaxed selection signature imply a shift in physiological role, which could affect endocrine sensitivity in trout populations exposed to xenobiotic estrogens. The study also highlights the importance of integrating MD‑derived thermodynamics with codon‑based selection analyses to infer the evolutionary fate of duplicated receptors.
Limitations and Future Directions
Only the LBD and a single ligand (E2) were examined; full‑length receptors, co‑activator interactions, and alternative ligands should be explored. Experimental validation (e.g., surface plasmon resonance, isothermal titration calorimetry) and tissue‑specific expression profiling would strengthen the computational conclusions.
Conclusion
Rainbow trout’s four ER isoforms display distinct ligand‑binding behaviors and evolutionary trajectories. ERα1 remains a high‑affinity, strongly conserved estrogen receptor, whereas ERα2 shows weakened binding and signs of relaxed selection, consistent with neofunctionalization. Both ERβ1 and ERβ2 retain similar affinities and are under uniform purifying selection, reflecting functional conservation. These findings illuminate how successive genome duplications can generate both preserved and diverged hormone‑receptor pathways in vertebrate lineages.
Comments & Academic Discussion
Loading comments...
Leave a Comment