Lymphotactin: how a protein can adopt two folds
Metamorphic proteins like Lymphotactin are a notable exception of the empirical principle that structured natural proteins possess a unique three dimensional structure. In particular, the human chemokine lymphotactin protein (Ltn) exists in two distinct conformations (one monomeric and one dimeric) under physiological conditions. In this work we use a Ca Go-model to show how this very peculiar behavior can be reproduced. From the study of the thermodynamics and of the kinetics we characterize the interconversion mechanism. In particular, this takes place through the docking of the two chains living in a third monomeric, partially unfolded, state which shows a residual structure involving a set of local contacts common to the two native conformations. The main feature of two-fold proteins appears to be the sharing of a common set of local contacts between the two distinct folds as confirmed by the study of two designed two-fold proteins. Metamorphic proteins may be more common than expected.
💡 Research Summary
The paper investigates the remarkable metamorphic behavior of the human chemokine lymphotactin (Ltn), which can exist simultaneously in two distinct three‑dimensional conformations under physiological conditions: a monomeric “Ltn‑10” fold and a dimeric “Ltn‑55” fold. Traditional protein science assumes a one‑sequence‑one‑structure relationship, but Ltn defies this rule, making it an ideal model for studying how a single amino‑acid chain can adopt multiple native states.
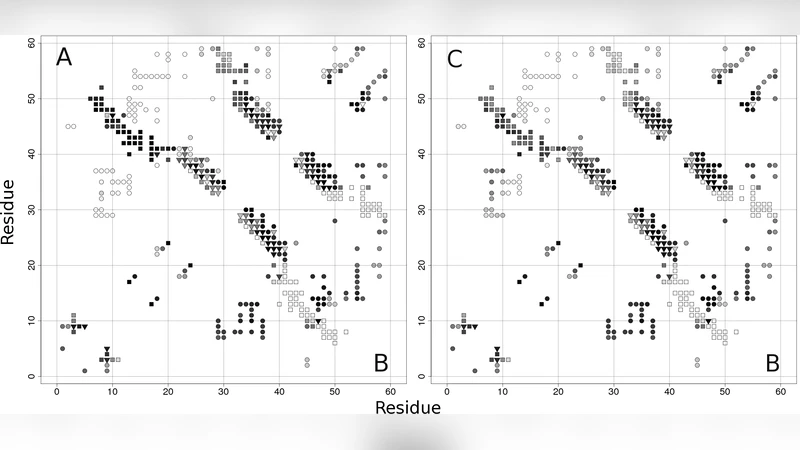
To explore this phenomenon, the authors construct a coarse‑grained Cα Go‑model that incorporates the native contact maps of both Ltn‑10 and Ltn‑55. By defining an energy function that rewards contacts present in either native structure, they generate a dual‑native energy landscape capable of stabilizing both folds. The model is calibrated against experimental data, confirming that the simulated temperature‑ and concentration‑dependent populations of the two states reproduce the observed equilibrium behavior.
Thermodynamic simulations reveal that at low temperature the monomeric fold is favored, at high temperature the dimeric fold dominates, and in an intermediate temperature window the free‑energy difference between the two states becomes vanishingly small. This narrow region corresponds to the experimentally reported coexistence of both conformations. Concentration scans further show that dimer formation scales with monomer concentration, indicating a concentration‑dependent docking step.
Kinetic analysis uncovers the actual interconversion pathway. Rather than a direct, large‑scale unfolding‑refolding event, the transition proceeds through a partially unfolded monomeric intermediate that retains a set of local contacts common to both native structures. These shared contacts are primarily short loops linking β‑strands and α‑helices. In the intermediate, the rest of the chain is largely disordered, allowing the two monomeric chains to approach each other. When they dock, the common contacts serve as nucleation points for re‑establishing the dimeric interface, while the previously unfolded regions refold to complete the Ltn‑55 structure. The calculated activation barrier is modest (≈2–3 kcal mol⁻¹), explaining why the conversion can occur rapidly under physiological conditions.
To test the generality of this mechanism, the authors design two artificial two‑fold proteins. Each engineered sequence contains distinct N‑ and C‑terminal structural elements but shares a core set of local contacts. Simulations demonstrate that these synthetic proteins also interconvert via a partially unfolded intermediate that preserves the shared contacts, confirming that the “common‑contact” principle is not unique to lymphotactin.
The study concludes that metamorphic proteins rely on a network of conserved local interactions that remain intact across different global folds. This network provides a scaffold that permits the protein to sample multiple native basins without complete loss of structure. Consequently, metamorphism may be far more prevalent in nature than previously thought, and the identified design principle offers a roadmap for engineering switchable proteins with tailored functional states. The findings have broad implications for protein design, drug discovery targeting conformational ensembles, and the understanding of diseases where mis‑folding and conformational switching play a role.
Comments & Academic Discussion
Loading comments...
Leave a Comment