Nonparametric Bayesian Density Modeling with Gaussian Processes
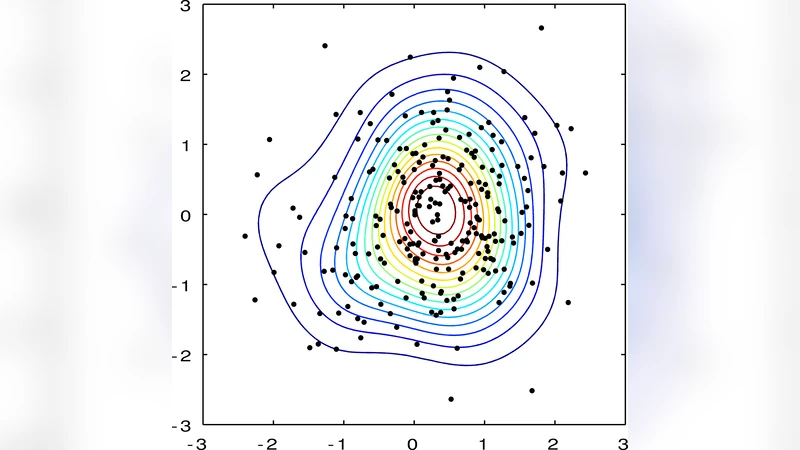
We present the Gaussian process density sampler (GPDS), an exchangeable generative model for use in nonparametric Bayesian density estimation. Samples drawn from the GPDS are consistent with exact, independent samples from a distribution defined by a density that is a transformation of a function drawn from a Gaussian process prior. Our formulation allows us to infer an unknown density from data using Markov chain Monte Carlo, which gives samples from the posterior distribution over density functions and from the predictive distribution on data space. We describe two such MCMC methods. Both methods also allow inference of the hyperparameters of the Gaussian process.
💡 Research Summary
The paper introduces the Gaussian Process Density Sampler (GPDS), a novel non‑parametric Bayesian model for density estimation that leverages the flexibility of Gaussian processes (GPs). Traditional non‑parametric approaches such as Dirichlet‑process mixtures or kernel density estimators either require explicit model‑order selection or suffer from the curse of dimensionality. GPDS circumvents these issues by defining a probability density directly from a GP draw. Concretely, a latent function f(x) is sampled from a GP prior with mean m(·) and covariance k(·,·). A logistic link σ(f)=1/(1+e^{‑f}) maps f to a positive function, and the density is obtained by normalizing this transformed function: p(x)=σ(f(x))/Z, where Z=∫σ(f(x))dx is the intractable normalizing constant. This construction guarantees exchangeability of the observed data and yields a prior over the entire space of continuous densities.
Because Z cannot be computed analytically, the authors develop two Markov chain Monte Carlo (MCMC) schemes to perform posterior inference over the latent function, the normalizing constant, and the GP hyper‑parameters. The first scheme augments the data with auxiliary variables u_i uniformly drawn from
Comments & Academic Discussion
Loading comments...
Leave a Comment