Early Detection of Breast Cancer using SVM Classifier Technique
This paper presents a tumor detection algorithm from mammogram. The proposed system focuses on the solution of two problems. One is how to detect tumors as suspicious regions with a very weak contrast to their background and another is how to extract features which categorize tumors. The tumor detection method follows the scheme of (a) mammogram enhancement. (b) The segmentation of the tumor area. (c) The extraction of features from the segmented tumor area. (d) The use of SVM classifier. The enhancement can be defined as conversion of the image quality to a better and more understandable level. The mammogram enhancement procedure includes filtering, top hat operation, DWT. Then the contrast stretching is used to increase the contrast of the image. The segmentation of mammogram images has been playing an important role to improve the detection and diagnosis of breast cancer. The most common segmentation method used is thresholding. The features are extracted from the segmented breast area. Next stage include, which classifies the regions using the SVM classifier. The method was tested on 75 mammographic images, from the mini-MIAS database. The methodology achieved a sensitivity of 88.75%.
💡 Research Summary
The paper proposes a computer‑aided detection (CAD) framework for early breast‑cancer identification using mammographic images from the mini‑MIAS database. The workflow consists of four sequential stages: (1) image enhancement, (2) tumor segmentation, (3) feature extraction, and (4) classification with a support vector machine (SVM).
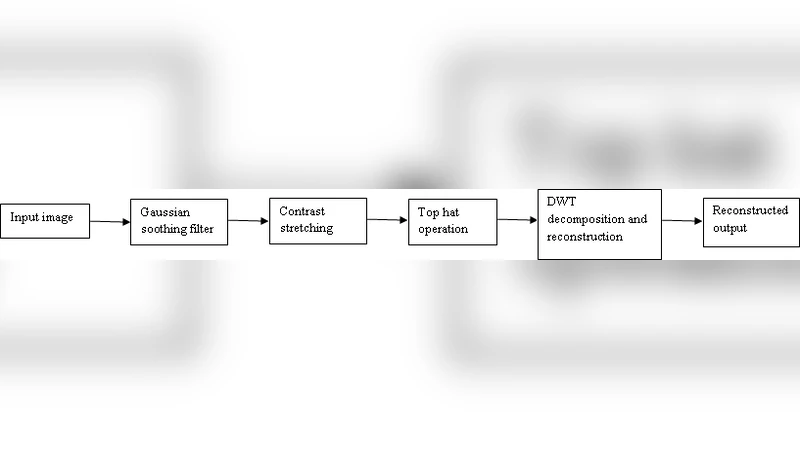
In the enhancement stage, the original gray‑scale mammogram is first smoothed with a Gaussian filter to suppress high‑frequency noise. A morphological top‑hat operation follows, employing a dark‑shaped structuring element to correct uneven illumination and to highlight bright structures (potential lesions) against a dark background. The top‑hat output is then decomposed into two scales using a discrete wavelet transform (DWT); the inverse DWT reconstruction yields an image in which tumor boundaries are more conspicuous.
Segmentation is performed by converting the enhanced image into binary masks through thresholding at several intensity levels. After thresholding, a second Gaussian smoothing pass removes isolated noisy pixels. The authors do not elaborate on how the optimal threshold(s) are selected, nor do they discuss alternative segmentation strategies such as region‑growing, active contours, or graph‑cut methods.
From each binary tumor region, ten morphological descriptors are computed: area, centroid, major axis length, minor axis length, eccentricity, orientation, filled area, extrema, solidity, and equivalent diameter. These features capture size and shape but ignore texture, intensity distribution, or higher‑order statistical information that are often discriminative for benign versus malignant lesions. No feature‑selection or dimensionality‑reduction procedure is reported.
The extracted feature vectors are fed into an SVM classifier. The manuscript outlines the basic SVM formulation, mentions four kernel families (linear, polynomial, radial basis function (RBF), and sigmoid), and presents the standard soft‑margin optimization problem. However, the experimental section fails to disclose which kernel was actually employed, the values of the regularization parameter C and kernel‑specific parameters (e.g., γ for RBF), or whether any hyper‑parameter tuning (grid search, cross‑validation) was performed. Consequently, the reported performance cannot be reproduced or fairly compared with other studies.
The authors evaluate the system on 75 mammograms, reporting a sensitivity of 88.75 %. No specificity, accuracy, precision, F1‑score, or area under the ROC curve (AUC) is provided, and the split between training and testing data is not described. With such a small dataset, the risk of overfitting is considerable, and the lack of a validation set makes it impossible to assess the model’s generalization capability.
In summary, the paper presents a conventional CAD pipeline that combines standard image‑preprocessing (Gaussian smoothing, top‑hat, DWT), simple global thresholding, a set of basic shape features, and an SVM classifier. While the approach yields a modest sensitivity on a limited test set, the study suffers from several methodological shortcomings: (i) insufficient justification and quantitative evaluation of the enhancement and segmentation steps; (ii) omission of texture or intensity‑based features that could improve discriminative power; (iii) lack of transparent SVM kernel selection and hyper‑parameter optimization; (iv) incomplete performance reporting (no specificity, ROC analysis); and (v) a small, non‑randomized dataset without proper cross‑validation.
Future work should address these gaps by (a) employing more robust segmentation techniques (e.g., multi‑scale level sets, deep‑learning‑based masks); (b) enriching the feature set with texture descriptors or deep features extracted from convolutional neural networks; (c) conducting systematic kernel and parameter selection using nested cross‑validation; (d) reporting a full suite of performance metrics and statistical significance tests; and (e) validating the method on larger, publicly available mammography collections (e.g., DDSM, INbreast) to demonstrate clinical relevance and reproducibility.
Comments & Academic Discussion
Loading comments...
Leave a Comment