Generalized Voronoi Tessellation as a Model of Two-dimensional Cell Tissue Dynamics
Voronoi tessellations have been used to model the geometric arrangement of cells in morphogenetic or cancerous tissues, however so far only with flat hypersurfaces as cell-cell contact borders. In order to reproduce the experimentally observed piecewise spherical boundary shapes, we develop a consistent theoretical framework of multiplicatively weighted distance functions, defining generalized finite Voronoi neighborhoods around cell bodies of varying radius, which serve as heterogeneous generators of the resulting model tissue. The interactions between cells are represented by adhesive and repelling force densities on the cell contact borders. In addition, protrusive locomotion forces are implemented along the cell boundaries at the tissue margin, and stochastic perturbations allow for non-deterministic motility effects. Simulations of the emerging system of stochastic differential equations for position and velocity of cell centers show the feasibility of this Voronoi method generating realistic cell shapes. In the limiting case of a single cell pair in brief contact, the dynamical nonlinear Ornstein-Uhlenbeck process is analytically investigated. In general, topologically distinct tissue conformations are observed, exhibiting stability on different time scales, and tissue coherence is quantified by suitable characteristics. Finally, an argument is derived pointing to a tradeoff in natural tissues between cell size heterogeneity and the extension of cellular lamellae.
💡 Research Summary
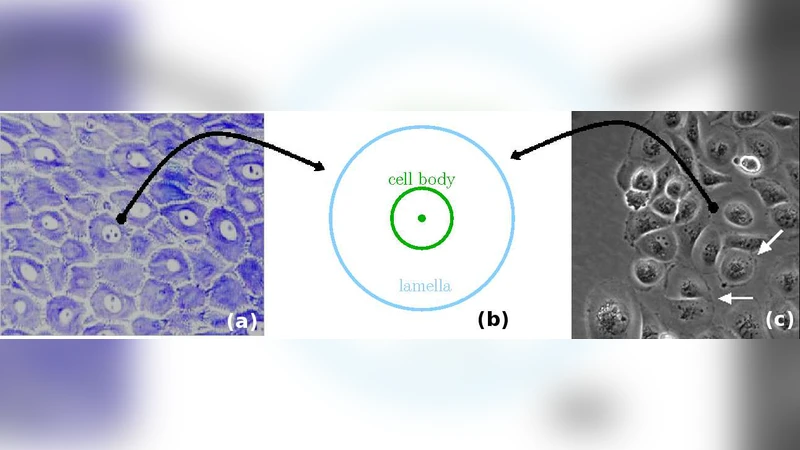
The paper presents a comprehensive framework for modeling two‑dimensional cell‑tissue dynamics using a generalized Voronoi tessellation based on multiplicatively weighted distance functions. Traditional Voronoi models assume flat cell‑cell interfaces, which fails to capture the piecewise‑spherical boundaries observed in real tissues. To overcome this limitation, each cell is represented by a circular “cell body” with center c_i and radius r_i. The distance from any point x to cell i is defined as d_i(x)=‖x−c_i‖/r_i, i.e., the Euclidean distance divided by the cell’s radius. This multiplicative weight naturally incorporates size heterogeneity: larger cells exert a weaker “effective distance” and therefore claim a larger region of space. The generalized Voronoi region V_i = {x | d_i(x) ≤ d_j(x) for all j ≠ i} is bounded by curved surfaces Σ_ij where d_i = d_j. These surfaces are spherical caps whose curvature depends on the ratio r_i/r_j, reproducing the experimentally observed rounded contact borders.
Cell‑cell mechanical interactions are modeled as surface force densities acting on each Σ_ij. Two contributions are considered: (1) an adhesive traction σ_a (negative sign) that pulls cells together when the interface distance falls below a threshold, and (2) a short‑range repulsive traction σ_r (positive sign) that sharply rises as cells approach each other, preventing overlap. The net force on cell i, F_i, is obtained by integrating the normal component of these densities over the entire boundary of V_i. This formulation yields forces that depend continuously on cell positions and radii, avoiding the discontinuities typical of classic vertex models.
At the tissue margin, cells generate protrusive locomotion forces, modeled as t_i n_i, where t_i is a scalar protrusive strength and n_i is the outward normal of the tissue edge. This term captures lamellipodial activity that drives tissue expansion, wound healing, or invasion.
The dynamics of each cell’s centre of mass obey a Newtonian equation with stochastic perturbations: m_i ẍ_i = F_i + ξ_i(t), where ξ_i(t) is a Gaussian white‑noise term with variance σ², representing non‑deterministic motility (e.g., cytoskeletal fluctuations). Consequently the system is a set of coupled stochastic differential equations (SDEs). Numerical integration is performed using the Euler‑Maruyama scheme with adaptive time stepping to resolve rapid force changes during cell‑cell contacts.
A special analytical focus is placed on the simplest configuration: a pair of cells in brief contact. By linearizing the interaction around the equilibrium separation, the inter‑cellular distance d(t) follows a nonlinear Ornstein‑Uhlenbeck (OU) process: ḋ = –k (d – d_eq) + σ η(t), where k is an effective spring constant derived from the combined adhesive/repulsive forces, d_eq is the equilibrium distance, and η(t) is standard Gaussian noise. This analysis yields closed‑form expressions for contact lifetime distributions and detachment probabilities, providing a benchmark for the full many‑cell simulations.
Extensive simulations explore a wide parameter space: cell‑size heterogeneity (distribution of r_i), protrusive strength t_i, adhesive/repulsive coefficients σ_a, σ_r, and noise amplitude σ. The emergent tissue morphologies display several key phenomena:
-
Size Heterogeneity Effects – Larger disparity in radii produces highly irregular Voronoi cells with pronounced curvature variations. The tissue’s topological connectivity (e.g., number of neighbours per cell) becomes broader, and local stress concentrations appear around very large or very small cells.
-
Lamellipodial Extension Trade‑off – Increasing t_i (longer lamellae) enhances tissue cohesion by enlarging contact areas, leading to smoother global shapes and slower rearrangements. However, excessive protrusion amplifies the magnitude of forces on the curved interfaces, making the system more susceptible to mechanical instability and spontaneous cell‑sorting events.
-
Stochastic Motility Influence – Higher noise levels σ accelerate cell rearrangements, promote occasional “fluid‑like” phases where cells exchange neighbours frequently, and can trigger a transition from a quasi‑solid packed state to a more fluidised configuration. This mirrors observations of epithelial tissues that fluidise during morphogenetic events.
From these observations the authors infer a biological trade‑off: natural tissues must balance cell‑size heterogeneity (which can be advantageous for functional specialization) against the need for sufficient lamellipodial reach to maintain mechanical integrity. Too much heterogeneity or overly extended lamellae each risk destabilising the tissue architecture.
The paper concludes by highlighting the model’s versatility. Because the weighted Voronoi construction directly yields cell shapes that can be compared with microscopy images, the framework can be calibrated against experimental data to infer mechanical parameters. Moreover, the stochastic formulation permits the study of processes where randomness plays a crucial role, such as tumor invasion, wound healing, and developmental patterning. Future extensions may incorporate three‑dimensional cell bodies, active contractility, or biochemical signaling fields, further bridging the gap between theoretical tissue mechanics and real biological systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment