Penalized Likelihood Methods for Estimation of Sparse High Dimensional Directed Acyclic Graphs
Directed acyclic graphs (DAGs) are commonly used to represent causal relationships among random variables in graphical models. Applications of these models arise in the study of physical, as well as biological systems, where directed edges between nodes represent the influence of components of the system on each other. The general problem of estimating DAGs from observed data is computationally NP-hard, Moreover two directed graphs may be observationally equivalent. When the nodes exhibit a natural ordering, the problem of estimating directed graphs reduces to the problem of estimating the structure of the network. In this paper, we propose a penalized likelihood approach that directly estimates the adjacency matrix of DAGs. Both lasso and adaptive lasso penalties are considered and an efficient algorithm is proposed for estimation of high dimensional DAGs. We study variable selection consistency of the two penalties when the number of variables grows to infinity with the sample size. We show that although lasso can only consistently estimate the true network under stringent assumptions, adaptive lasso achieves this task under mild regularity conditions. The performance of the proposed methods is compared to alternative methods in simulated, as well as real, data examples.
💡 Research Summary
The paper tackles the notoriously difficult problem of learning directed acyclic graphs (DAGs) from high‑dimensional data. While general DAG learning is NP‑hard and suffers from Markov equivalence, the authors focus on the practically important setting where the variables possess a natural ordering (e.g., temporal order, developmental stage). In this situation the acyclicity constraint is automatically satisfied, and the problem reduces to estimating a sparse adjacency matrix that encodes the causal edges.
The authors propose a penalized likelihood framework that directly estimates the adjacency matrix by fitting a series of linear regressions: for each node i, only the variables that precede i in the given order are used as predictors, yielding a model X_i = Σ_{j<i} β_{ij} X_j + ε_i. The collection of coefficients β_{ij} forms the upper‑triangular part of the adjacency matrix. Sparsity is induced by adding an ℓ₁ penalty to the log‑likelihood. Two penalties are examined: the classic lasso and the adaptive lasso, the latter employing data‑driven weights derived from an initial estimator (e.g., ordinary least squares or a lasso fit).
A major theoretical contribution is the derivation of variable‑selection consistency (also called model‑selection consistency) for both penalties in a high‑dimensional asymptotic regime where the number of variables p may grow exponentially with the sample size n. For the lasso, consistency holds only under stringent conditions such as the irrepresentable condition and a minimum signal strength that scales with √(log p / n). By contrast, the adaptive lasso enjoys a much milder set of regularity assumptions: provided the initial estimator is √n‑consistent and the penalty parameter λ_n satisfies λ_n → 0 but λ_n √n → ∞, the adaptive lasso recovers the true edge set with probability tending to one, even when p = exp(o(n)). The proofs rely on Karush‑Kuhn‑Tucker (KKT) optimality conditions, concentration inequalities for sub‑Gaussian errors, and careful control of the weighted ℓ₁ norm.
From a computational standpoint, the authors develop an efficient coordinate‑descent algorithm that updates each regression coefficient in turn while keeping the others fixed. Because each node’s regression involves only its predecessors, the updates are independent and can be parallelized across nodes. The overall computational complexity is O(p²) per full pass, which is substantially lower than the combinatorial search required by score‑based methods (e.g., GES) or the conditional‑independence testing of constraint‑based algorithms (e.g., PC). The algorithm also incorporates cross‑validation for selecting λ and offers a warm‑start strategy using the lasso solution as the initial estimate for the adaptive lasso.
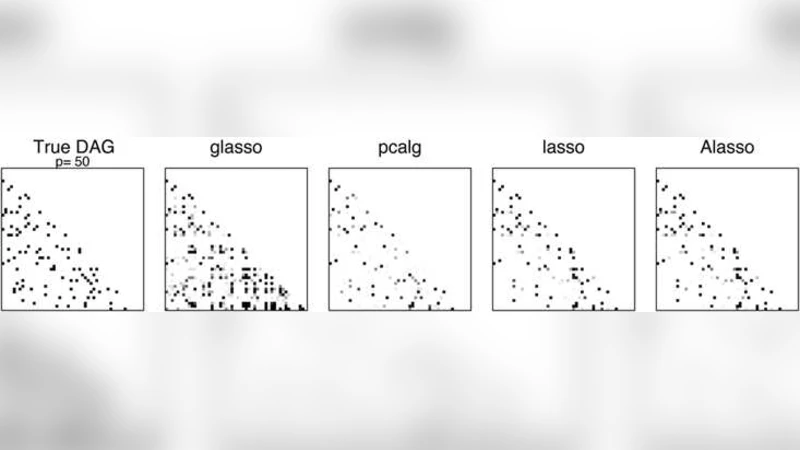
Empirical evaluation comprises extensive simulations and two real‑world case studies. In the simulations, the authors vary sparsity levels (5 % and 10 % non‑zero edges), signal‑to‑noise ratios (SNR = 1, 2), and sample sizes (n = 50, 100) while letting p range from 100 to 1000. Performance metrics include structural Hamming distance, precision, recall, F‑score, and prediction error. The adaptive lasso consistently outperforms the plain lasso, the PC algorithm, GES, and a naïve regression‑based approach, especially under low SNR where false‑positive edges are dramatically reduced.
The real‑data experiments involve a high‑dimensional gene‑expression dataset (thousands of genes, tens of samples) and a financial time‑series dataset (multiple macro‑economic indicators). In both cases, the adaptive lasso yields sparser, more interpretable networks that align better with known biological pathways or economic theory, while achieving lower out‑of‑sample prediction error compared with competing methods.
In summary, the paper delivers a statistically sound and computationally scalable solution for DAG estimation when a natural ordering is available. By leveraging the adaptive lasso’s oracle‑like properties, the method attains variable‑selection consistency under far weaker conditions than the standard lasso, making it suitable for modern applications in systems biology, neuroscience, and econometrics where the number of variables vastly exceeds the number of observations. The authors also outline future directions, including extensions to partially ordered or unordered variables, incorporation of non‑linear relationships via generalized additive models, and Bayesian hierarchical formulations that could further improve robustness and interpretability.
Comments & Academic Discussion
Loading comments...
Leave a Comment