On the basic computational structure of gene regulatory networks
Gene regulatory networks constitute the first layer of the cellular computation for cell adaptation and surveillance. In these webs, a set of causal relations is built up from thousands of interactions between transcription factors and their target genes. The large size of these webs and their entangled nature make difficult to achieve a global view of their internal organisation. Here, this problem has been addressed through a comparative study for {\em Escherichia coli}, {\em Bacillus subtilis} and {\em Saccharomyces cerevisiae} gene regulatory networks. We extract the minimal core of causal relations, uncovering the hierarchical and modular organisation from a novel dynamical/causal perspective. Our results reveal a marked top-down hierarchy containing several small dynamical modules for \textit{E. coli} and \textit{B. subtilis}. Conversely, the yeast network displays a single but large dynamical module in the middle of a bow-tie structure. We found that these dynamical modules capture the relevant wiring among both common and organism-specific biological functions such as transcription initiation, metabolic control, signal transduction, response to stress, sporulation and cell cycle. Functional and topological results suggest that two fundamentally different forms of logic organisation may have evolved in bacteria and yeast.
💡 Research Summary
This study tackles the challenge of deciphering the global organization of large‑scale gene regulatory networks (GRNs) by performing a comparative analysis of three model organisms: Escherichia coli, Bacillus subtilis, and the yeast Saccharomyces cerevisiae. The authors first assemble high‑quality, directed GRNs from curated databases (RegulonDB, DBTBS, YEASTRACT), removing self‑loops and duplicate edges, resulting in networks comprising 2–6 K genes and 5–15 K regulatory interactions.
To extract the essential computational backbone, they introduce the concept of a “causal minimal core.” Using a greedy algorithm that evaluates each edge’s contribution to preserving all input‑to‑output causal paths and overall network robustness, they iteratively prune edges while monitoring connectivity. The resulting core retains only about 8–12 % of the original edges yet contains the majority of information‑flow pathways and feedback loops, effectively representing the network’s computational skeleton.
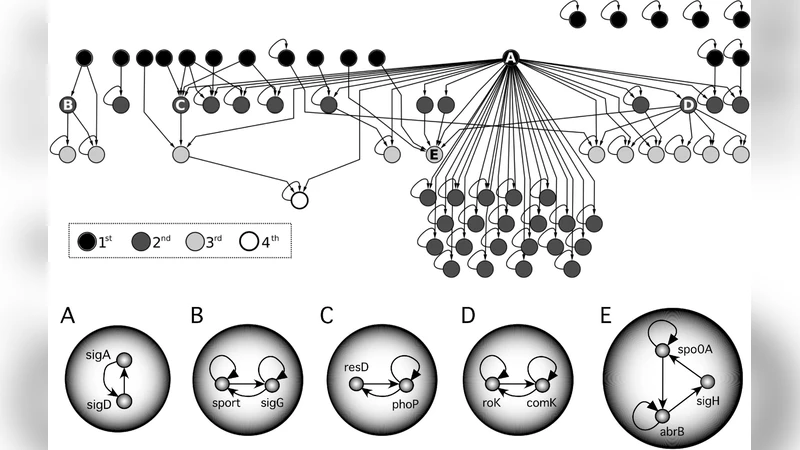
The core is then decomposed into hierarchical layers through topological sorting, yielding an upstream “input” layer (predominantly transcription factors), a downstream “output” layer (target genes involved in metabolism, stress response, etc.), and one or more intermediate layers. Within these layers, the authors identify “dynamic modules” – tightly interconnected subgraphs that exhibit autonomous dynamical behaviors such as switches, oscillators, or multi‑stable states when simulated with Boolean or ODE‑based models.
In E. coli and B. subtilis, the hierarchy is pronounced: a small number of high‑level TFs feed into three to five modest dynamic modules, each comprising roughly 30–50 genes. These modules are sparsely linked, often through single or double feedback connections, producing a distributed, top‑down architecture that enables rapid, localized responses to environmental cues. Functional enrichment shows these modules are specialized for transcription initiation, amino‑acid biosynthesis, oxidative‑stress mitigation, and sporulation.
Yeast displays a markedly different organization. Its core collapses into a classic “bow‑tie” architecture: numerous upstream signals converge onto a single, large dynamic module occupying the network’s centre. This module contains over 300 genes and integrates pathways governing the cell cycle, DNA repair, central carbon metabolism, and signal transduction. The internal wiring is dense, with multiple overlapping feedback and feed‑forward loops, allowing complex dynamical regimes (e.g., sustained oscillations, bistability) that simultaneously process diverse inputs and generate coordinated outputs. Downstream, the information disperses to many specialized gene sets, reflecting a centralized, integrative logic.
The authors validate these structural observations with two complementary analyses. First, Gene Ontology and KEGG pathway enrichment maps each module to biological processes, confirming that bacterial modules are functionally narrow while the yeast central module is functionally broad. Second, robustness tests involving random node/edge removal and targeted deletion of core TFs reveal contrasting resilience patterns: bacterial networks collapse sharply when a single module is disrupted, whereas yeast retains substantial functionality due to redundant pathways within its massive central module.
Key insights emerging from the work are: (1) a systematic method for distilling large GRNs into a minimal causal core that preserves computational essence; (2) identification of two fundamentally different logic architectures—distributed small‑module hierarchies in prokaryotes versus a centralized large‑module bow‑tie in eukaryotes; (3) a link between these architectures and evolutionary pressures, with bacteria favoring rapid, modular adaptability and yeast favoring integrated control of complex cellular programs.
The study’s implications extend to synthetic biology and systems medicine. By mimicking the bacterial modular hierarchy, engineered circuits can achieve fast, fault‑tolerant responses; by emulating the yeast bow‑tie, designers can build compact yet multifunctional synthetic modules capable of coordinating diverse cellular tasks. Future directions proposed include extending the causal‑core framework to higher eukaryotes, quantitatively modeling the dynamics of identified modules, and probing how environmental fluctuations reshape module composition over evolutionary timescales. Overall, the paper provides a novel dynamical‑causal lens for understanding how cells compute, offering both conceptual advances and practical tools for network engineering.
Comments & Academic Discussion
Loading comments...
Leave a Comment