Statistical physics of pairwise probability models
Statistical models for describing the probability distribution over the states of biological systems are commonly used for dimensional reduction. Among these models, pairwise models are very attractive in part because they can be fit using a reasonable amount of data: knowledge of the means and correlations between pairs of elements in the system is sufficient. Not surprisingly, then, using pairwise models for studying neural data has been the focus of many studies in recent years. In this paper, we describe how tools from statistical physics can be employed for studying and using pairwise models. We build on our previous work on the subject and study the relation between different methods for fitting these models and evaluating their quality. In particular, using data from simulated cortical networks we study how the quality of various approximate methods for inferring the parameters in a pairwise model depends on the time bin chosen for binning the data. We also study the effect of the size of the time bin on the model quality itself, again using simulated data. We show that using finer time bins increases the quality of the pairwise model. We offer new ways of deriving the expressions reported in our previous work for assessing the quality of pairwise models.
💡 Research Summary
The paper investigates how tools from statistical physics can be used to study and apply pairwise probability models, which are widely employed for dimensional reduction of biological systems, especially neural data. A pairwise model assumes that the probability distribution over a set of binary variables (e.g., spike/no‑spike) can be captured solely by the first‑order moments (means) and second‑order moments (pairwise correlations). Because only these low‑order statistics are required, the model can be fitted with relatively modest amounts of data, making it attractive for large neural populations.
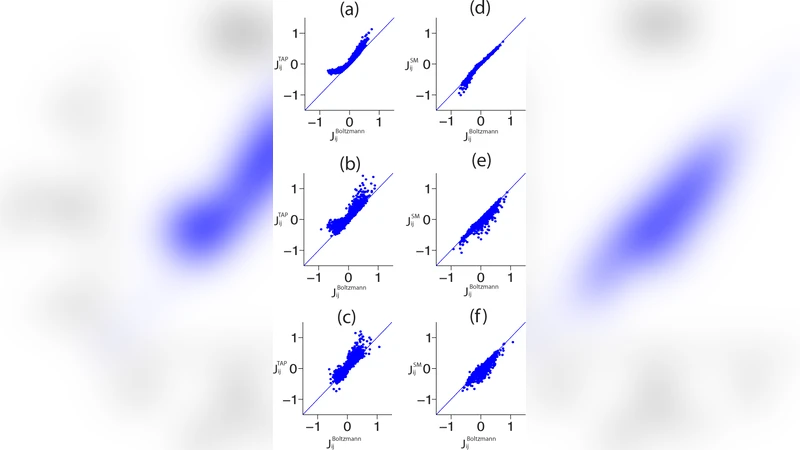
The authors first review several common approximate inference methods for estimating the model parameters (the fields h_i and couplings J_ij). The naïve mean‑field (MF) approach linearizes the inverse problem, treating the correlation matrix as if the variables were weakly coupled. The Thouless‑Anderson‑Palmer (TAP) expansion adds second‑order corrections, incorporating higher‑order cumulants and thus improving accuracy at the cost of more complex self‑consistency equations. The pseudo‑likelihood (PL) method maximizes the product of conditional probabilities for each variable given the rest, dramatically reducing computational cost while retaining good statistical efficiency. The paper clarifies the theoretical assumptions underlying each method and discusses their computational scaling.
To evaluate these methods, the authors generate synthetic spike trains from a biologically realistic cortical network model. The simulated network contains thousands of spiking neurons with heterogeneous connectivity and external drive, providing a ground‑truth set of parameters for a known pairwise model. The continuous spike times are discretized into binary time bins of various widths (1 ms, 5 ms, 20 ms, 50 ms). For each bin size, the empirical means ⟨s_i⟩ and pairwise correlations ⟨s_i s_j⟩ are computed, and the three inference algorithms are applied to recover the fields and couplings.
The quality of the recovered models is quantified using two complementary metrics. First, the Kullback‑Leibler (KL) divergence between the true distribution (known from the simulation) and the inferred distribution measures how well the full probability landscape is captured. Second, the difference in log‑likelihood (or equivalently the entropy gap) evaluates how much predictive power is lost when using the approximate model. Across all bin sizes, the PL method consistently yields the smallest KL divergence and entropy gap, closely matching the ground‑truth parameters. TAP performs better than MF for intermediate bin widths but suffers from convergence issues when the coupling matrix becomes strongly non‑diagonal. MF, while computationally cheapest, shows substantial bias especially when the bin is large and the effective correlations are strong.
A central finding is the systematic dependence of model quality on the bin size. Finer bins (≤ 5 ms) preserve the temporal precision of spikes, leading to accurate estimates of both means and correlations. Consequently, the inferred pairwise model reproduces the true distribution with high fidelity. Coarser bins aggregate multiple spikes into a single binary event, effectively smoothing out high‑frequency fluctuations and attenuating strong, nonlinear interactions. This loss of information manifests as inflated KL divergences and larger entropy gaps, particularly for MF and TAP. The authors therefore recommend using the smallest feasible bin that still yields reliable statistics, balancing temporal resolution against the need for sufficient sample counts per bin.
Beyond empirical evaluation, the paper contributes a new derivation of the quality‑assessment formulas previously introduced by the authors. By employing a cumulant expansion and diagrammatic techniques, they separate the contributions of first‑order (mean) and second‑order (pairwise) cumulants from higher‑order terms. This formalism clarifies why MF fails when higher‑order cumulants are non‑negligible and why TAP’s second‑order correction improves performance only up to a point. The new derivation also yields explicit error bounds that depend on the magnitude of the couplings and the bin size, offering a theoretical tool for predicting when a given approximation will be reliable.
In summary, the study provides a comprehensive comparison of three widely used inference schemes for pairwise models, demonstrates how temporal discretization critically influences model fidelity, and offers a refined theoretical framework for assessing model quality. The results have practical implications for experimental design: when applying pairwise maximum‑entropy models to neural recordings, researchers should aim for the finest temporal bin compatible with their data volume, and they should prefer pseudo‑likelihood estimation for its robustness and scalability. Moreover, the analytical error bounds derived from the cumulant expansion can guide the choice of approximation method in other domains where pairwise models are employed, such as genomics, protein‑interaction networks, and social‑behavioral data.
Comments & Academic Discussion
Loading comments...
Leave a Comment