Scaling, phase transition and genus distribution functions in matrix models of RNA with linear external interactions
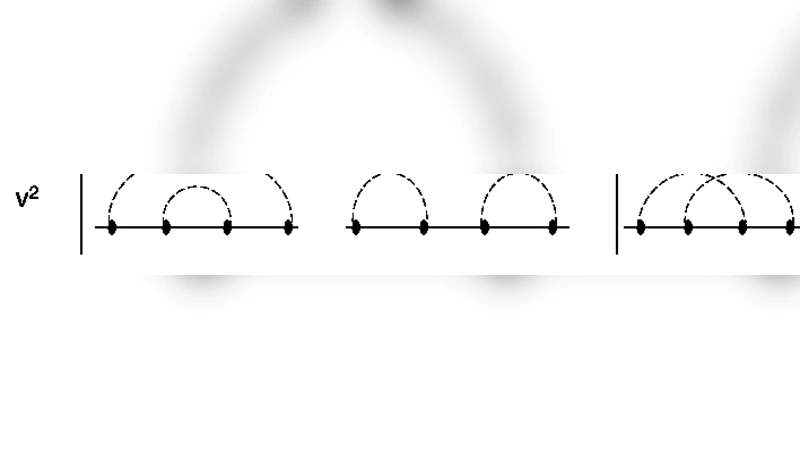
A linear external perturbation is introduced in the action of the partition function of the random matrix model of RNA [G. Vernizzi, H. Orland and A. Zee, Phys. Rev. Lett. 94, 168103 (2005)]. It is seen that (i). the perturbation distinguishes between paired and unpaired bases in that there are structural changes, from unpaired and paired base structures ($0 \leq \alpha < 1$) to completely paired base structures ($\alpha=1$), as the perturbation parameter $\alpha$ approaches 1 ($\alpha$ is the ratio of interaction strengths of original and perturbed terms in the action of the partition function), (ii). the genus distributions exhibit small differences for small even and odd lengths $L$, (iii). the partition function of the linear interacting matrix model is related via a scaling formula to the re-scaled partition function of the random matrix model of RNA, (iv). the free energy and specific heat are plotted as functions of $L$, $\alpha$ and temperature $T$ and their first derivative with respect to $\alpha$ is plotted as a function of $\alpha$. The free energy shows a phase transition at $\alpha=1$ for odd (both small and large) lengths and for even lengths the transition at $\alpha=1$ gets sharper and sharper as more pseudoknots are included (that is for large lengths).
💡 Research Summary
The paper extends the random‑matrix model of RNA secondary structure by adding a linear external perturbation to the action. In the original model the interaction term is quartic (∝ Tr M⁴) and generates the combinatorial weight for base‑pairing; the new term α Tr M is linear in the matrix and is controlled by a dimensionless parameter α that measures the ratio of the external interaction strength to the original one. When α = 0 the model reduces to the standard one, allowing both paired and unpaired bases. As α increases, the linear term increasingly favours configurations in which every nucleotide participates in a pair, and at α = 1 the model forces a completely paired structure.
Mathematically the partition function for a sequence of length L is written as
\
Comments & Academic Discussion
Loading comments...
Leave a Comment