Seeing Science
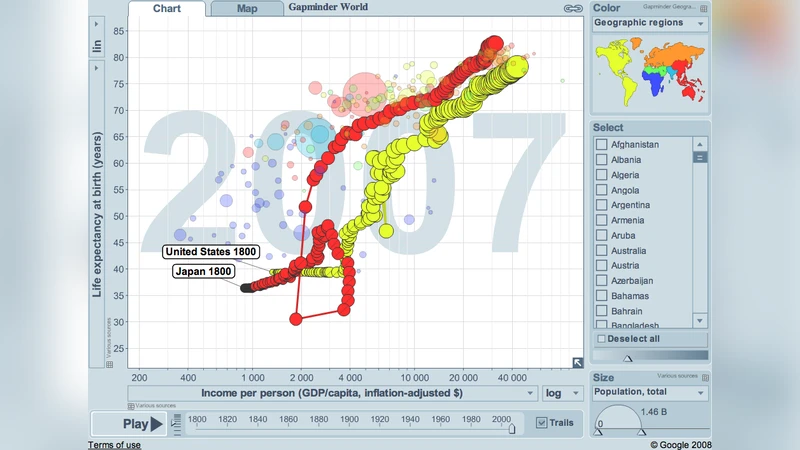
The ability to represent scientific data and concepts visually is becoming increasingly important due to the unprecedented exponential growth of computational power during the present digital age. The data sets and simulations scientists in all fields can now create are literally thousands of times as large as those created just 20 years ago. Historically successful methods for data visualization can, and should, be applied to today’s huge data sets, but new approaches, also enabled by technology, are needed as well. Increasingly, “modular craftsmanship” will be applied, as relevant functionality from the graphically and technically best tools for a job are combined as-needed, without low-level programming.
💡 Research Summary
The paper “Seeing Science” addresses the growing challenge of visualizing ever‑larger scientific data sets in the digital age and proposes a hybrid strategy that blends time‑tested visualization techniques with modern, modular toolchains. The authors begin by documenting the exponential increase in data volume and complexity across disciplines, noting that traditional static plots and simple 2‑D graphics are insufficient for conveying the richness of contemporary simulations, high‑resolution observations, and multi‑parameter experiments. They review historically successful visualizations—such as Maxwell’s field line renderings, Newtonian orbital diagrams, and GIS‑based geographic maps—to illustrate how well‑designed visual metaphors can align with human perceptual strengths and the underlying structure of the data.
The central contribution of the paper is the concept of “modular craftsmanship.” Rather than building monolithic, custom‑coded visualization pipelines, researchers are encouraged to assemble a suite of specialized, interoperable modules that each excel at a particular function: high‑performance graphics engines (e.g., Unity, Unreal) for real‑time 3‑D rendering, interactive dashboard frameworks (e.g., Plotly Dash, Bokeh) for web‑based exploration, machine‑learning dimensionality‑reduction tools (e.g., t‑SNE, UMAP) for compressing high‑dimensional data, and cloud‑based streaming/rendering services (e.g., AWS, Azure) for handling massive data flows. By exposing well‑defined APIs and plug‑in interfaces, these modules can be combined without low‑level programming, allowing scientists to tailor a visualization pipeline that matches the specific demands of their data.
To guide module selection, the authors introduce a “Visualization Design Matrix” that evaluates four axes: data dimensionality, temporal dynamics, required level of user interaction, and visual fidelity. For example, a climate‑modeling project with multi‑petabyte, time‑varying fields would pair a GPU‑accelerated volume renderer with a slider‑based UI, while a neuro‑connectomics study would integrate a graph database (Neo4j) with a WebGL network visualizer to explore complex connectivity patterns.
Three case studies demonstrate the practical impact of this approach. In the first, climate scientists combined a GPU‑driven 3‑D volume renderer with an interactive web dashboard, enabling simultaneous inspection of temperature, precipitation, and wind fields across the globe. In the second, neuroscientists visualized whole‑brain connectomes by linking a graph database to a Three.js‑based interactive network viewer, revealing previously hidden hub structures. In the third, a physics laboratory streamed real‑time experimental data through an AWS Kinesis pipeline into a WebGL viewer, providing immediate feedback that reduced iteration cycles.
The discussion acknowledges potential pitfalls: mismatched interfaces between modules can create integration overhead; performance bottlenecks may arise when individually optimized components are combined; and licensing incompatibilities between open‑source and commercial tools can hinder distribution. To mitigate these issues, the authors recommend adopting a common API schema, containerizing each module with Docker/Kubernetes for reproducible deployment, and establishing community‑driven standards for visualization interoperability.
In conclusion, the paper argues that visual representation remains a cornerstone of scientific discovery and communication. By embracing modular craftsmanship, researchers can build flexible, reusable, and scalable visualization pipelines that keep pace with data growth, foster interdisciplinary collaboration, and enhance transparency. The authors also propose regular “visualization workshops” to train scientists in assembling and customizing module stacks, thereby cultivating a culture where seeing data is as essential as analyzing it. This paradigm shift promises to transform how science is explored, validated, and shared with both expert and public audiences.
Comments & Academic Discussion
Loading comments...
Leave a Comment