Physico-chemical modelling of target depletion during hybridisation on oligonulceotide microarrays
The effect of target molecule depletion from the supernatant solution is incorporated into a physico-chemical model of hybridisation on oligonucleotide microarrays. Two possible regimes are identified: local depletion, in which depletion by a given probe feature only affects that particular probe, and global depletion, in which all features responding to a given target species are affected. Examples are given of two existing spike-in data sets experiencing measurable effects of target depletion. The first of these, from an experiment by Suzuki et al. using custom built arrays with a broad range of probe lengths and mismatch positions, is verified to exhibit local and not global depletion. The second dataset, the well known Affymetrix HGU133a latin square experiment is shown to be very well explained by a global depletion model. It is shown that microarray calibrations relying on Langmuir isotherm models which ignore depletion effects will significantly underestimate specific target concentrations. It is also shown that a combined analysis of perfect match and mismatch probe signals in terms of a simple graphical summary, namely the hook curve method, can discriminate between cases of local and global depletion.
💡 Research Summary
The paper addresses a frequently overlooked source of bias in microarray hybridisation experiments: the depletion of target molecules from the hybridisation solution as they bind to surface‑immobilised probes. Traditional analyses assume that the concentration of free target remains essentially constant, allowing the use of a simple Langmuir isotherm to relate probe intensity to target concentration. The authors argue that this assumption breaks down when the amount of target bound to the array is non‑negligible compared with the total amount present in the solution. To capture this effect they extend the Langmuir model by introducing a depletion term and develop two distinct regimes.
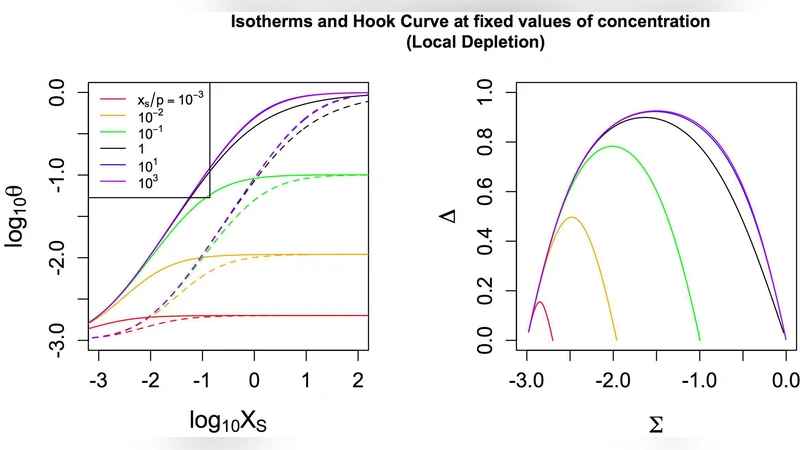
In the “local depletion” regime each probe feature is treated as an isolated micro‑reactor; the binding of target to a particular probe reduces the local free concentration only for that feature. Consequently, each probe’s signal can be described by its own binding constant and a local depletion parameter, without influencing neighbouring features. In the “global depletion” regime all probes that recognise the same target share a common solution pool. Binding to any of these probes lowers the overall free concentration, thereby affecting the binding equilibrium of every other probe for that target. This leads to a coupled system of equations in which a single global depletion parameter governs the entire set of probes.
The authors validate both models using two publicly available spike‑in data sets. The first, from Suzuki et al., employed custom arrays with a wide variety of probe lengths and mismatch positions. Because each probe was effectively isolated, the data are best described by the local depletion model; fitting a global depletion model yields large systematic errors, especially at low target concentrations where the predicted intensities saturate prematurely. The second data set is the classic Affymetrix HGU133a Latin‑square experiment, in which dozens of perfect‑match (PM) and mismatch (MM) probes interrogate the same set of spiked‑in transcripts. Here the global depletion model provides an excellent fit, reproducing the characteristic non‑linear relationship between PM and MM intensities, the early onset of saturation, and the concentration‑dependent curvature observed across the Latin‑square concentration levels.
A further contribution is the use of the “hook curve” graphical summary, which plots PM intensity against MM intensity for each probe set. Under ideal Langmuir behaviour the points trace a symmetric hook‑shaped curve. The authors show that local depletion preserves this symmetry, whereas global depletion distorts the curve, producing an asymmetric hook with a pronounced bend at high concentrations. This visual diagnostic allows researchers to quickly assess whether depletion effects are local or global without performing full non‑linear fitting.
Finally, the paper quantifies the impact of ignoring depletion on concentration estimates. Simulations and fits to the experimental data reveal that neglecting the depletion term can lead to systematic under‑estimation of target concentrations by 20–40 % when the depletion parameter lies in the range 0.1–0.3. Such errors are especially problematic for low‑abundance transcripts, clinical biomarker assays, and any application that relies on accurate absolute quantification. The authors therefore recommend incorporating depletion‑aware models into standard microarray analysis pipelines and using the hook‑curve diagnostic as a routine quality‑control step. This work provides both a theoretical framework and practical tools to improve the reliability of microarray‑based measurements in the presence of target depletion.
Comments & Academic Discussion
Loading comments...
Leave a Comment