📝 Original Info
- Title: Residue network in protein native structure belongs to the universality class of three dimensional critical percolation cluster
- ArXiv ID: 0809.4876
- Date: 2009-11-13
- Authors: Researchers from original ArXiv paper
📝 Abstract
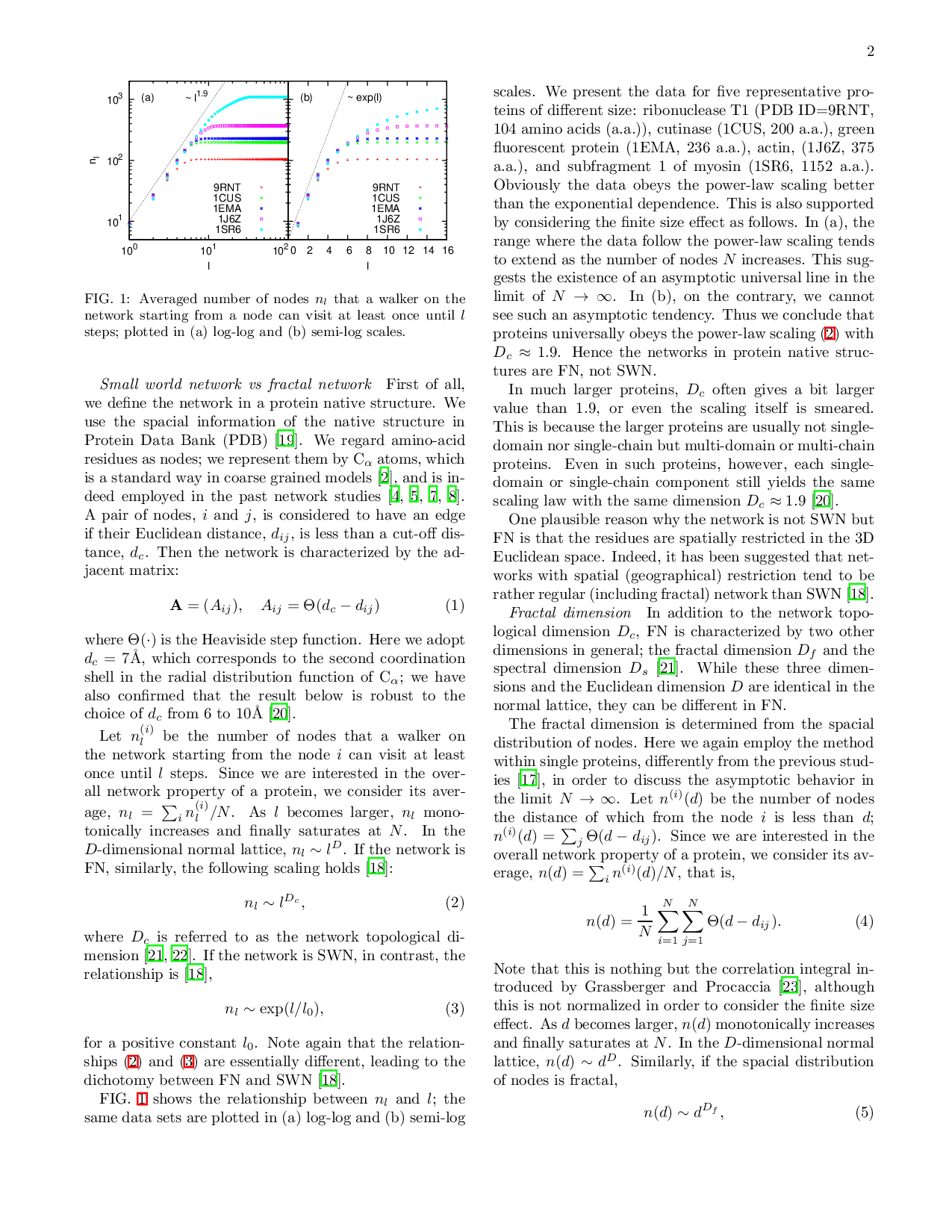
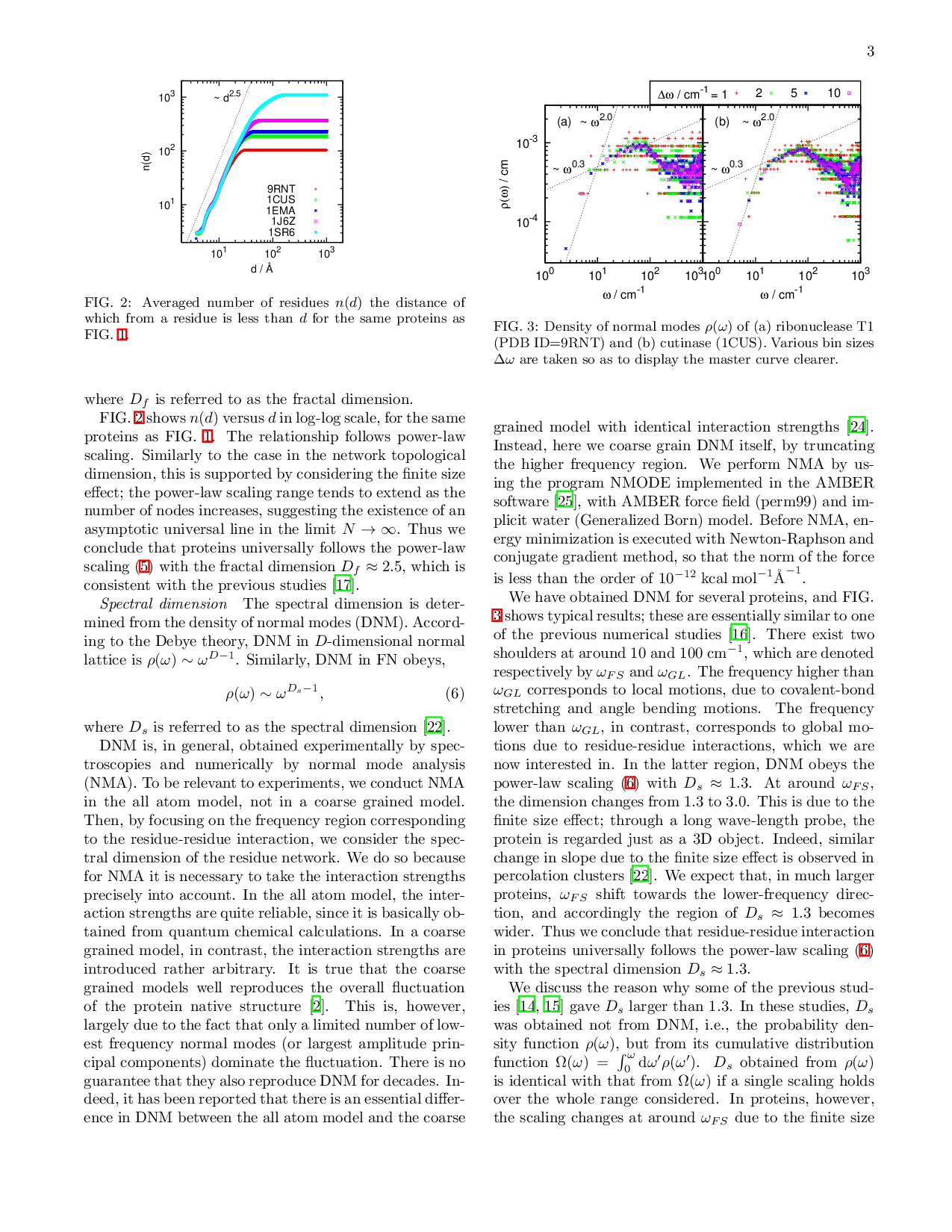
A single protein molecule is regarded as a contact network of amino-acid residues. Some studies have indicated that this network is a small world network (SWN), while other results have implied that this is a fractal network (FN). However, SWN and FN are essentially different in the dependence of the shortest path length on the number of nodes. In this paper, we investigate this dependence in the residue contact networks of proteins in native structures, and show that the networks are not SWN but FN. FN is generally characterized by several dimensions. Among them, we focus on three dimensions; the network topological dimension $D_c$, the fractal dimension $D_f$, and the spectral dimension $D_s$. We find that proteins universally yield $D_c \approx 1.9$, $D_f \approx 2.5$ and $Ds \approx 1.3$. These values are in surprisingly good coincidence with those in three dimensional critical percolation cluster. Hence the residue contact networks in the protein native structures belong to the universality class of three dimensional percolation cluster. The criticality is relevant to the ambivalent nature of the protein native structures, i.e., the coexistence of stability and instability, both of which are necessary for a protein to function as a molecular machine or an allosteric enzyme.
💡 Deep Analysis
Deep Dive into Residue network in protein native structure belongs to the universality class of three dimensional critical percolation cluster.
A single protein molecule is regarded as a contact network of amino-acid residues. Some studies have indicated that this network is a small world network (SWN), while other results have implied that this is a fractal network (FN). However, SWN and FN are essentially different in the dependence of the shortest path length on the number of nodes. In this paper, we investigate this dependence in the residue contact networks of proteins in native structures, and show that the networks are not SWN but FN. FN is generally characterized by several dimensions. Among them, we focus on three dimensions; the network topological dimension $D_c$, the fractal dimension $D_f$, and the spectral dimension $D_s$. We find that proteins universally yield $D_c \approx 1.9$, $D_f \approx 2.5$ and $Ds \approx 1.3$. These values are in surprisingly good coincidence with those in three dimensional critical percolation cluster. Hence the residue contact networks in the protein native structures belong to the un
📄 Full Content
arXiv:0809.4876v1 [physics.bio-ph] 28 Sep 2008
Residue network in protein native structure belongs to
the universality class of three dimensional critical percolation cluster
Hidetoshi Morita∗and Mitsunori Takano†
Faculty of Science and Engineering, Waseda University, Tokyo 169-8555, Japan
(Dated: June 14, 2021)
A single protein molecule is regarded as a contact network of amino-acid residues. Some studies
have indicated that this network is a small world network (SWN), while other results have implied
that this is a fractal network (FN). However, SWN and FN are essentially different in the dependence
of the shortest path length on the number of nodes. In this paper, we investigate this dependence
in the residue contact networks of proteins in native structures, and show that the networks are
not SWN but FN. FN is generally characterized by several dimensions. Among them, we focus
on three dimensions; the network topological dimension Dc, the fractal dimension Df, and the
spectral dimension Ds. We find that proteins universally yield Dc ≈1.9, Df ≈2.5 and Ds ≈1.3.
These values are in surprisingly good coincidence with those in three dimensional critical percolation
cluster. Hence the residue contact networks in the protein native structures belong to the universality
class of three dimensional percolation cluster. The criticality is relevant to the ambivalent nature
of the protein native structures, i.e., the coexistence of stability and instability, both of which are
necessary for a protein to function as a molecular machine or an allosteric enzyme.
PACS numbers: 87.15.-V, 64.60.-i, 89.75.He, 05.45.Df
Introduction
Proteins are one-dimensional chains of
amino-acid residues embedded in three dimensional (D =
3; 3D) Euclidean space.
The residues neighboring in
the Euclidean space are in contact with each other.
Thus we can regard a protein molecule as a contact net-
work of amino-acid residues [1, 2]. This network view-
point is complementary to the energy landscape picture
[3] in understanding the general properties of proteins.
We hereafter consider this network within single protein
molecules in their native structures, in particular focus-
ing on its universality among proteins.
Some recent studies [4, 5, 6, 7, 8] have applied the lat-
est network theory to the residue network, by regarding
the amino-acid residues and their contacts as nodes and
edges, respectively. The important quantities to charac-
terize the network are the clustering coefficient C and the
shortest path length L [9]. Those studies have demon-
strated that in the residue networks C is larger than the
random networks [10] while L is smaller than the normal
lattice. This indicates that the residue network is a small
world network (SWN) [11].
On the other hand, the spacial profile of residues within
single protein molecules has long been studied with the
use of authentic methods of material science.
Earlier
spectroscopic studies [12] have shown anomalous density
of states.
These results, accompanied with theoretical
studies [13], have suggested that the protein structures
possess the property of fractal lattice.
The fractality
within single proteins has also been supported numeri-
cally through the density of normal modes [14, 15, 16]
and the spacial mass distribution [17]. This implies that
the residue network that we are interested in is a fractal
network (FN).
From the general viewpoint of the network theory,
however, there lies a dichotomy between SWN and
FN [18].
The clustering coefficient C cannot discrimi-
nate between SWN and FN, since in both the networks
C have a larger value than the random networks. In con-
trast, the dependence of the shortest path length L on
the number of nodes N is essentially different between
SWN and FN; L depends on N logarithmically and alge-
braically, respectively. By exploiting the N-dependency
of L, we can differentiate SWN and FN, in principle.
In proteins, nevertheless, it is practically difficult to
clearly distinguish between these two N-dependence.
This is because the size of proteins does not distribute
widely enough to cover sufficient decades. The same data
sets can be read as a straight line both in log-log (SWN)
and semi-log (FN) plot.
To overcome this difficulty, here we introduce a more
sophisticated method. Instead of the N-L plot among
various sized proteins, we investigate an equivalent within
single protein molecules; we calculate the number of
nodes nl that can be reached until l path steps. Then, by
overdrawing the nl-l plot for various sized proteins, we
obtain a universal curve, as well as the deviation from it
due to finite size effect. Thus we can discuss an asymp-
totic behavior in the large N limit. We thereby find that
network in protein native structures is FN, not SWN.
This is the first result of this letter.
We then obtain the three characteristic dimensions of
fractal residue network; the network topological dimen-
sion Dc, the fractal dimension Df, and the spectral di-
mension Ds.
The v
…(Full text truncated)…
📸 Image Gallery
Reference
This content is AI-processed based on ArXiv data.