Emergence of species in evolutionary simulated annealing
Which factors govern the evolution of mutation rates and emergence of species? Here, we address this question using a first principles model of life where population dynamics of asexual organisms is coupled to molecular properties and interactions of proteins encoded in their genomes. Simulating evolution of populations, we found that fitness increases in punctuated steps via epistatic events, leading to formation of stable and functionally interacting proteins. At low mutation rates, species - populations of organisms with identical genotypes - form, while at higher mutation rates, species are lost through delocalization in sequence space without an apparent loss of fitness. However, when mutation rate was a selectable trait, the population initially maintained high mutation rate until a high fitness level is reached, after which organisms with low mutation rates are gradually selected, with the population eventually reaching mutation rates comparable to those of modern DNA-based organisms. These results provide microscopic insights into the dynamic fitness landscape of asexual populations of unicellular organisms.
💡 Research Summary
The paper presents a first‑principles computational framework that couples population dynamics of asexual unicellular organisms to the molecular properties of the proteins they encode. Each organism’s genome specifies a set of proteins; for each protein the model assigns a folding free energy (ΔG_folding) and, when applicable, a binding free energy (ΔG_binding) with other proteins. The overall fitness of an organism is defined as a function of the stability of all its proteins and the strength of the interaction network they form. By explicitly incorporating epistatic effects—where a single mutation can simultaneously improve the folding of several proteins and strengthen multiple protein‑protein contacts—the authors are able to map a rugged, high‑dimensional fitness landscape that evolves over time.
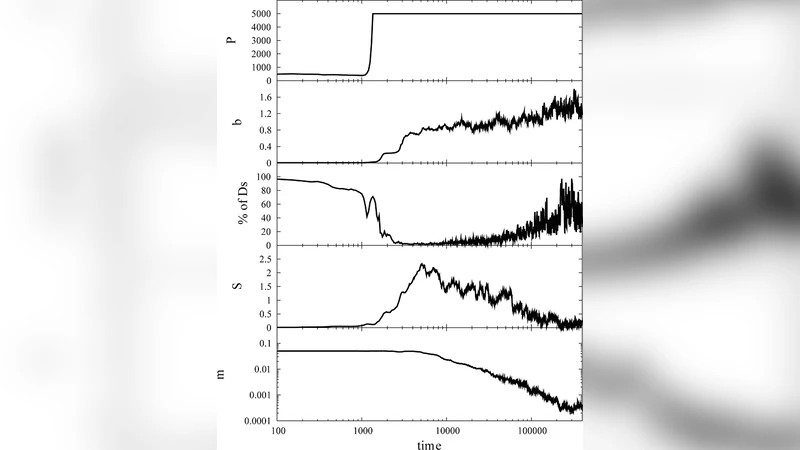
Two major simulation regimes are explored. In the first, the mutation rate (μ) is held constant while random point mutations accumulate. Fitness does not increase smoothly; instead it rises in punctuated steps. Each step corresponds to an epistatic event that creates a new, more stable configuration of the interaction network, effectively moving the population to a higher fitness “peak.” This behavior mirrors the concept of punctuated equilibrium and demonstrates that large fitness gains can arise from rare, cooperative changes rather than the gradual accumulation of independent beneficial mutations.
In the second regime, μ itself is treated as an evolvable trait. Initially, a high mutation rate (≈10⁻⁴ per gene per generation) is favored because it generates extensive genetic diversity, allowing the population to explore the fitness landscape rapidly and achieve a high fitness plateau. Once the plateau is reached, the cost of accumulating deleterious mutations (mutational load) becomes significant. Consequently, genotypes with lower μ begin to outcompete high‑μ genotypes, and the average mutation rate drifts downward. The simulation converges on mutation rates in the range 10⁻⁸–10⁻⁹, which closely matches measured rates in modern DNA‑based microbes. This dynamic illustrates a natural trade‑off between adaptability (high μ) and genomic integrity (low μ) and provides a mechanistic explanation for why contemporary organisms maintain such low mutation rates despite the evolutionary advantage of occasional rapid adaptation.
A key conceptual contribution is the operational definition of a “species” as a sub‑population of organisms sharing an identical genotype. Under low mutation rates, the population clusters tightly in sequence space, forming discrete genotype clusters that correspond to classical species. At higher mutation rates, the genotype distribution spreads (delocalization), erasing clear species boundaries even though the overall fitness of the population remains high. This finding suggests that species cohesion is not strictly required for maintaining high fitness in asexual populations; rather, it is a by‑product of low mutational flux.
Parameter choices—protein length, distribution of ΔG_folding and ΔG_binding, and the statistical properties of mutational effects—are calibrated against empirical protein stability data and observed mutational spectra. The authors demonstrate that the model reproduces realistic evolutionary dynamics, including the emergence of low‑mutation‑rate regimes comparable to those of extant microbes.
In summary, the study delivers three major insights: (1) mutation rate is a non‑linear driver of both evolutionary speed and species formation; (2) epistatic events can generate punctuated fitness improvements by reorganizing the protein interaction network; and (3) when mutation rate is selectable, evolution initially exploits high mutational variability to climb the fitness landscape, then gradually selects for low mutational load, arriving at mutation rates characteristic of modern life. These results deepen our microscopic understanding of asexual evolution, provide a quantitative bridge between molecular biophysics and population genetics, and have practical implications for synthetic biology, directed evolution, and the design of more efficient evolutionary algorithms.
Comments & Academic Discussion
Loading comments...
Leave a Comment