Identifying short motifs by means of extreme value analysis
The problem of detecting a binding site – a substring of DNA where transcription factors attach – on a long DNA sequence requires the recognition of a small pattern in a large background. For short binding sites, the matching probability can display large fluctuations from one putative binding site to another. Here we use a self-consistent statistical procedure that accounts correctly for the large deviations of the matching probability to predict the location of short binding sites. We apply it in two distinct situations: (a) the detection of the binding sites for three specific transcription factors on a set of 134 estrogen-regulated genes; (b) the identification, in a set of 138 possible transcription factors, of the ones binding a specific set of nine genes. In both instances, experimental findings are reproduced (when available) and the number of false positives is significantly reduced with respect to the other methods commonly employed.
💡 Research Summary
The paper addresses a fundamental statistical challenge in the detection of short transcription‑factor binding sites (TFBS) within long DNA promoters. Conventional motif‑finding pipelines typically (i) model the background and the motif using Markov chains, (ii) compute a log‑likelihood score Wₖ for each possible window of length ℓ, and (iii) assume that the distribution of these scores is Gaussian, thereby applying standard significance tests. This Gaussian assumption breaks down for short motifs (ℓ ≈ 5–20) because the score is a sum of only a few independent nucleotide contributions, so the central‑limit theorem does not apply. The authors demonstrate, using artificial Position‑Specific Frequency Matrices (PSFMs) on simulated promoters, that the fraction of score distributions failing a normality test can exceed 30 % for short motifs, indicating that the extreme‑value distribution of the maximum score is rarely the Gumbel law expected for Gaussian variables.
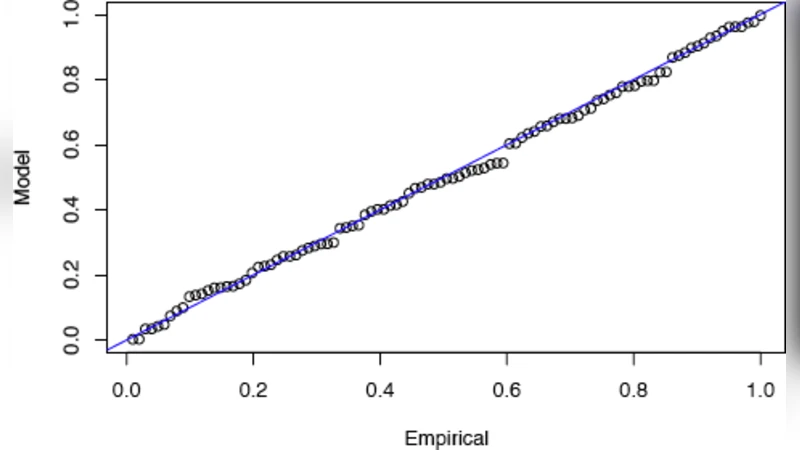
To overcome this limitation, the authors adopt extreme‑value theory, specifically the Peak‑over‑Threshold (POT) method. After computing the full set of scores {Wₖ} for a promoter, they select a high threshold u and examine the excesses Yₖ = Wₖ − u that exceed u. Theory (Pickands‑Balkema‑de Haan) tells us that, for sufficiently high u, the excesses follow a Generalized Pareto Distribution (GPD) characterized by a shape parameter ξ and a scale parameter σ. The authors estimate ξ and σ by fitting the empirical excess distribution, using the mean‑excess plot eₙ(u) versus u to locate a region where eₙ(u) is approximately linear (eₙ(u) ≈ σ + ξ u/(1 − ξ)). This provides an objective way to choose u that balances bias (u must be high enough for the GPD approximation to hold) and variance (enough exceedances must remain for reliable parameter estimation).
Once ξ and σ are obtained, the number N of exceedances above u is modeled as a Poisson variable with rate λ = N/(L − ℓ + 1), where L is the promoter length. For each exceedance Yₖ, the probability that a larger exceedance occurs somewhere in the promoter is
Pₖ = 1 − exp
Comments & Academic Discussion
Loading comments...
Leave a Comment