Controlling Viral Capsid Assembly with Templating
We develop coarse-grained models that describe the dynamic encapsidation of functionalized nanoparticles by viral capsid proteins. We find that some forms of cooperative interactions between protein subunits and nanoparticles can dramatically enhance rates and robustness of assembly, as compared to the spontaneous assembly of subunits into empty capsids. For large core-subunit interactions, subunits adsorb onto core surfaces en masse in a disordered manner, and then undergo a cooperative rearrangement into an ordered capsid structure. These assembly pathways are unlike any identified for empty capsid formation. Our models can be directly applied to recent experiments in which viral capsid proteins assemble around the functionalized inorganic nanoparticles [Sun et al., Proc. Natl. Acad. Sci (2007) 104, 1354]. In addition, we discuss broader implications for understanding the dynamic encapsidation of single-stranded genomic molecules during viral replication and for developing multicomponent nanostructured materials.
💡 Research Summary
The paper presents a coarse‑grained computational framework for studying the dynamic encapsidation of functionalized inorganic nanoparticles by viral capsid proteins. By representing each capsid subunit as a rigid body with directional interaction sites and modeling the nanoparticle core as a spherical surface bearing tunable attractive sites, the authors independently vary two key parameters: the core‑subunit binding energy (ε_core) and the subunit‑subunit binding energy (ε_pp). Systematic simulations across a wide range of ε_core values reveal two distinct assembly regimes.
In the weak‑core regime (low ε_core), subunits first nucleate on the nanoparticle surface, forming a small ordered patch that then grows by sequential addition of individual subunits. This pathway closely mirrors the well‑characterized nucleation‑and‑growth mechanism of empty capsid formation, and it proceeds relatively slowly. The resulting capsids are often incomplete or malformed because the surface coverage is low and the nucleation event is stochastic.
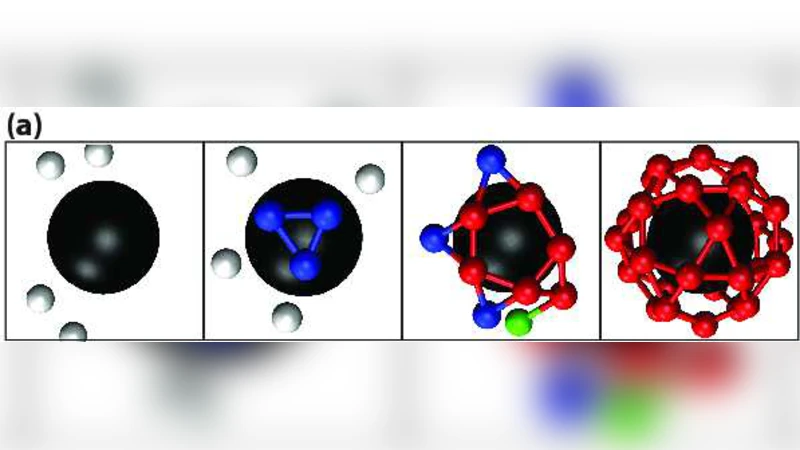
When ε_core exceeds a critical threshold, a qualitatively different pathway emerges. Subunits adsorb en masse onto the core in a disordered fashion, creating a densely packed but amorphous layer. Because the core provides a geometric template and a strong attractive field, the entire layer undergoes a cooperative rearrangement: subunits collectively reorient, break and reform inter‑subunit contacts, and finally lock into the icosahedral symmetry of a mature capsid. This collective transition is rapid, yields near‑perfect capsids, and is far more robust to fluctuations in subunit concentration or temperature. The authors term this mechanism “mass adsorption followed by cooperative rearrangement,” and they demonstrate that it can accelerate assembly by an order of magnitude compared with the nucleation‑driven route.
The model reproduces experimental observations reported by Sun et al. (PNAS 2007), where capsid proteins encapsulated gold nanoparticles functionalized with various ligands. In those experiments, increasing the surface charge density of the particles dramatically improved encapsidation efficiency and speed, exactly as predicted by increasing ε_core in the simulations. Additional parameter sweeps show that particle radius, surface charge density, and ligand spacing modulate the balance between the two pathways: small, highly charged particles favor the cooperative rearrangement route, whereas larger, weakly charged particles tend to follow the traditional nucleation pathway.
Beyond reproducing these experiments, the study offers broader biological insight. The cooperative rearrangement mechanism provides a plausible explanation for how single‑stranded viral genomes, which are highly negatively charged, can act as “soft templates” that attract capsid proteins en masse and then guide them into an ordered capsid without a distinct nucleation event. This suggests that genome‑mediated templating may be a general strategy employed by many RNA viruses to achieve rapid and reliable assembly inside host cells.
Finally, the authors discuss implications for nanomaterial design. By engineering nanoparticle surfaces (e.g., ligand chemistry, charge patterning, curvature), one can deliberately invoke the cooperative rearrangement pathway to produce uniform, protein‑coated nanostructures. Such hybrid particles could serve as drug delivery vehicles, catalytic scaffolds, or components of bio‑electronic devices. The paper thus bridges virology, soft‑matter physics, and nanotechnology, providing both a mechanistic understanding of templated capsid assembly and a practical roadmap for exploiting this phenomenon in engineered systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment