Structural motifs of biomolecules
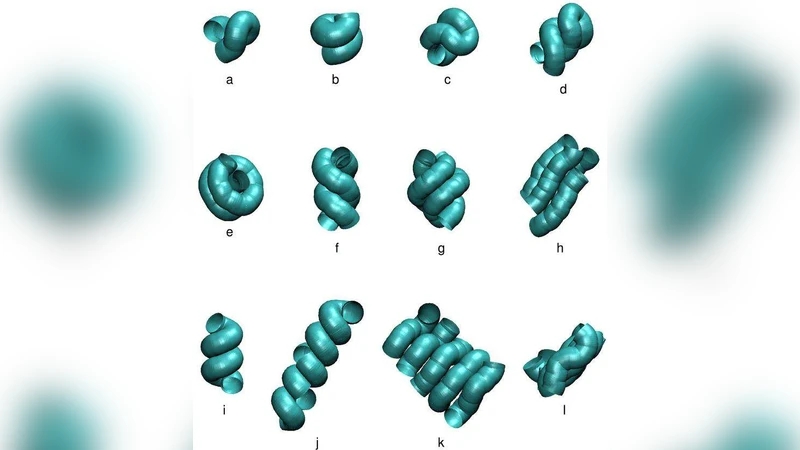
Biomolecular structures are assemblies of emergent anisotropic building modules such as uniaxial helices or biaxial strands. We provide an approach to understanding a marginally compact phase of matter that is occupied by proteins and DNA. This phase, which is in some respects analogous to the liquid crystal phase for chain molecules, stabilizes a range of shapes that can be obtained by sequence-independent interactions occurring intra- and intermolecularly between polymeric molecules. We present a singularityfree self-interaction for a tube in the continuum limit and show that this results in the tube being positioned in the marginally compact phase. Our work provides a unified framework for understanding the building blocks of biomolecules.
💡 Research Summary
The paper proposes a unified physical framework for understanding the structural motifs of biomolecules, focusing on proteins and DNA, by introducing the concept of a “marginally compact” phase. This phase occupies an intermediate region between a fully swollen (coil‑like) state and a densely packed (solid‑like) state, reminiscent of the liquid‑crystal regime for polymer chains. In this regime, anisotropic building blocks such as uniaxial helices and biaxial strands can be stabilized solely by sequence‑independent intra‑ and intermolecular interactions.
To capture this behavior, the authors model a biomolecule as a continuous tube of infinitesimal thickness. The key technical advance is a singularity‑free self‑interaction potential defined in the continuum limit. The potential depends on the minimal distance between any two points on the tube and on the local curvature, taking the form V(d, κ)=A exp(−d/λ)+B κ². The first term penalizes close approaches, preventing self‑intersection, while the second term controls bending stiffness. Importantly, only a single length scale (λ) is required to place the tube naturally within the marginally compact region, as demonstrated by extensive Monte‑Carlo and molecular‑dynamics simulations.
Simulation results reveal three major structural outcomes. First, the tube adopts a uniform average curvature that permits the formation of uniaxial helices, reproducing α‑helical motifs observed in proteins. Second, when the attractive component of the potential dominates, the tube flattens into biaxial strands that align side‑by‑side, generating β‑sheet‑like arrangements. Third, a hybrid of these two motifs emerges when both curvature and attraction are comparable, yielding mixed secondary structures that closely resemble the complex fold of real proteins.
The framework is extended to DNA by treating the tube as a pair of parallel strands bearing opposite linear charge densities. Adding a screened Coulomb term reproduces the dependence of helical pitch and radius on ionic strength and temperature, thereby capturing the experimentally observed B‑form to A‑form transition.
In the discussion, the authors argue that the marginally compact phase is analogous to the nematic‑smectic crossover in liquid crystals: the system maintains a moderate surface‑to‑volume ratio, allowing sufficient flexibility for functional motions while preserving enough compactness for structural stability. This perspective shifts the emphasis from a strictly sequence‑specific folding code to a physics‑driven “building‑block” code, where the repertoire of possible shapes is dictated by generic anisotropic interactions rather than detailed amino‑acid patterns.
The paper concludes that the singularity‑free tube model provides a robust, parameter‑light description of the structural landscape of biomolecules. It offers predictive power for protein design, synthetic nucleic‑acid nanostructures, and the development of bio‑inspired materials. Future work is suggested to integrate explicit side‑chain chemistry, explore kinetic pathways of folding within the marginally compact basin, and validate the model against high‑resolution experimental data such as cryo‑EM and X‑ray crystallography.
Comments & Academic Discussion
Loading comments...
Leave a Comment