An evolutionary model with Turing machines
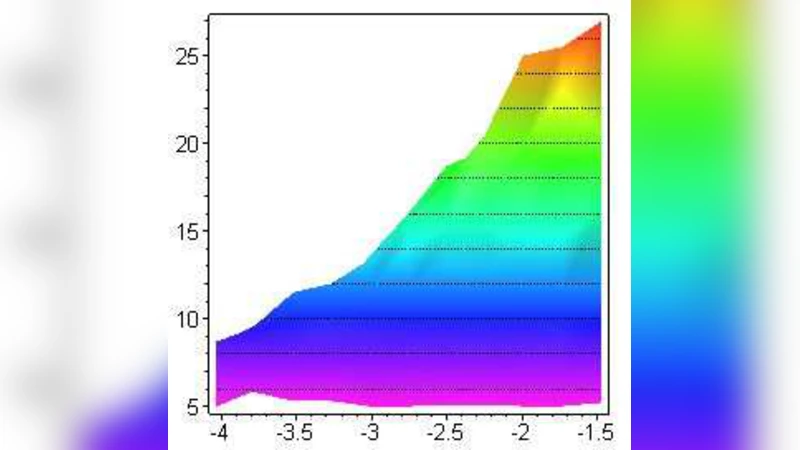
The development of a large non-coding fraction in eukaryotic DNA and the phenomenon of the code-bloat in the field of evolutionary computations show a striking similarity. This seems to suggest that (in the presence of mechanisms of code growth) the evolution of a complex code can’t be attained without maintaining a large inactive fraction. To test this hypothesis we performed computer simulations of an evolutionary toy model for Turing machines, studying the relations among fitness and coding/non-coding ratio while varying mutation and code growth rates. The results suggest that, in our model, having a large reservoir of non-coding states constitutes a great (long term) evolutionary advantage.
💡 Research Summary
The paper investigates a striking parallel between two phenomena that appear in very different domains: the large proportion of non‑coding DNA in eukaryotic genomes and the “code‑bloat” that frequently emerges in evolutionary‑computation systems. Both involve the accumulation of large inactive portions of a program or genome while the system is still capable of evolving functional complexity. The authors ask whether a substantial reservoir of inactive material is actually advantageous when mechanisms for code growth are present.
To explore this hypothesis they construct a toy evolutionary model based on Turing machines (TMs). In the model each individual is a TM defined by a finite set of states and a transition table. “Coding” states are those that are visited during the execution of a given input; “non‑coding” states are never activated and therefore have no immediate effect on the output. The fitness of an individual is measured by how closely the TM’s output for a set of predefined inputs matches a target output string; exact matches receive high scores, mismatches are penalized.
Two evolutionary operators are applied each generation. The first is a mutation operator that randomly alters existing transitions or inserts/deletes transitions, thereby changing the functional part of the machine. The second is a code‑growth operator that adds new states that are initially non‑coding, mimicking the insertion of transposable elements or other neutral DNA in real genomes. The rates of these two processes are controlled independently: μ denotes the mutation rate and γ the code‑growth rate. By varying μ and γ across a wide range, the authors generate many evolutionary runs that span thousands of generations.
The results reveal several consistent patterns. When μ is low, fitness improves only modestly regardless of how many non‑coding states are present; the population quickly reaches a plateau because beneficial mutations are rare. When μ is high, however, populations that maintain a large pool of non‑coding states (high non‑coding fraction) are far more likely to discover beneficial new transitions. The non‑coding pool acts as a genetic buffer: it absorbs the disruptive effects of many mutations while providing raw material that can later be co‑opted into functional code. In other words, neutral or “junk” states increase the mutational robustness of the system and expand the searchable space for innovation.
The code‑growth rate γ also shows a non‑linear effect. Small γ values lead to insufficient non‑coding material, limiting the long‑term adaptability of the population. Very large γ values cause an explosion of inactive states, which raises memory and computational costs and eventually slows down the evolutionary process. An intermediate γ yields the best trade‑off: enough neutral states to buffer mutations and to be recruited later, but not so many that the system becomes computationally unwieldy.
Long‑term simulations (several thousand generations) demonstrate that populations with a non‑coding fraction above roughly 70 % are markedly more successful at evolving complex target functions, such as multi‑step logical operations. Populations with a non‑coding fraction below about 30 % may achieve high fitness early on, but they tend to stagnate or even regress when the task requires additional layers of complexity.
These findings have two major implications. First, in evolutionary algorithms the ubiquitous code‑bloat should not be dismissed merely as inefficiency; deliberately allowing or even encouraging the growth of neutral code can improve the algorithm’s ability to explore rugged fitness landscapes and to discover novel solutions. Second, the study provides quantitative support for the biological view that non‑coding DNA is not merely evolutionary “junk” but a reservoir of latent functionality that can be harnessed during periods of rapid environmental change or increased mutational pressure.
The authors conclude by suggesting future work that would endow non‑coding states with richer structural properties (e.g., repetitive motifs, transposable‑element‑like dynamics) and by comparing the model’s predictions with empirical genomic data. Overall, the paper offers a unified conceptual framework that links code growth, non‑coding reservoirs, and evolvability, delivering insights valuable to both the evolutionary‑computation community and to researchers studying genome evolution.
Comments & Academic Discussion
Loading comments...
Leave a Comment