Length control of microtubules by depolymerizing motor proteins
In many intracellular processes, the length distribution of microtubules is controlled by depolymerizing motor proteins. Experiments have shown that, following non-specific binding to the surface of a microtubule, depolymerizers are transported to the microtubule tip(s) by diffusion or directed walk and, then, depolymerize the microtubule from the tip(s) after accumulating there. We develop a quantitative model to study the depolymerizing action of such a generic motor protein, and its possible effects on the length distribution of microtubules. We show that, when the motor protein concentration in solution exceeds a critical value, a steady state is reached where the length distribution is, in general, non-monotonic with a single peak. However, for highly processive motors and large motor densities, this distribution effectively becomes an exponential decay. Our findings suggest that such motor proteins may be selectively used by the cell to ensure precise control of MT lengths. The model is also used to analyze experimental observations of motor-induced depolymerization.
💡 Research Summary
The paper presents a quantitative theoretical framework for understanding how depolymerizing motor proteins regulate the length distribution of microtubules (MTs) inside cells. The authors begin by reviewing experimental observations that members of the kinesin‑13 family (e.g., MCAK, Kif2a) bind nonspecifically to the MT lattice, travel to the plus or minus end either by diffusion or by directed, ATP‑driven walks, and then catalyze rapid depolymerization once they accumulate at the tip. While these phenomena are well documented, a unified mathematical description linking motor concentration, transport dynamics, and the resulting steady‑state MT length distribution has been lacking.
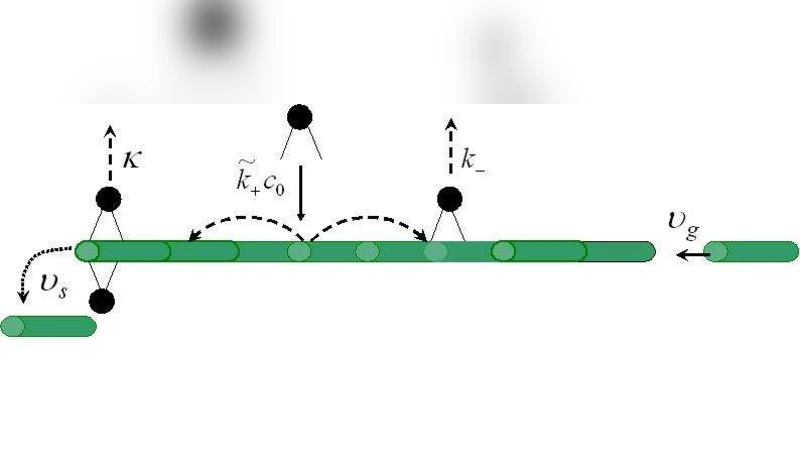
To fill this gap, the authors construct a one‑dimensional continuum model of an MT of length L. Motor proteins in solution at bulk concentration C_bulk bind to the lattice with rate k_on and detach with rate k_off. Once bound, a motor either diffuses with coefficient D or walks with velocity v toward the tip; the model can accommodate a mixture of both processes by superposition. The flux of motors arriving at the tip, J(L), is derived from the steady‑state solution of the reaction‑diffusion‑advection equation. At the tip, motors accumulate and generate a depolymerization rate proportional to the number of tip‑localized motors, n_tip, i.e., k_dep·n_tip, with an upper bound k_dep^max that captures the saturation of the depolymerizing activity for highly processive motors. The tip dynamics obey dn_tip/dt = J(L) – k_dep·n_tip, while the MT length changes according to dL/dt = –k_dep·n_tip.
Analysis of the coupled equations reveals two distinct regimes determined by the bulk motor concentration relative to a critical value C_crit. When C_bulk < C_crit, the tip flux is insufficient to sustain appreciable depolymerization; MTs either keep growing or display a monotonic, long‑tailed length distribution that diverges in the mean. In this “unstable” regime the system lacks a well‑defined steady state.
When C_bulk > C_crit, a stable steady state emerges. The length distribution becomes non‑monotonic, featuring a single peak at a characteristic length L*. This peak arises because the arrival time of motors at the tip, τ_arrival ≈ L²/D for diffusion‑dominated transport or τ_arrival ≈ L/v for directed walks, sets a length‑dependent feedback: longer MTs receive motors more slowly, allowing them to persist, whereas shorter MTs are rapidly depolymerized. The position of L* shifts toward shorter lengths as D or v increase, or as k_on rises, reflecting faster motor delivery.
A particularly interesting limit is that of highly processive motors (very low k_off). In this case, tip accumulation quickly reaches the saturation limit n_sat = k_dep^max/k_dep, and the depolymerization rate becomes effectively constant. The resulting length distribution collapses to an exponential decay, indistinguishable from a simple first‑order decay process. This finding contrasts with the classic “dynamic instability” picture, where catastrophe and rescue events dominate length control; here, motor‑mediated depolymerization alone can generate exponential length statistics.
The theoretical predictions are confronted with in‑vitro experiments using purified kinesin‑13 proteins. By varying motor concentration and measuring MT shrinkage rates and final length histograms, the authors demonstrate quantitative agreement with the model: (1) a clear threshold concentration above which MT lengths become narrowly distributed, (2) a shift of the distribution peak with changes in ATP concentration (which modulates v), and (3) an exponential tail emerging at high motor densities. Parameter fitting yields realistic values for k_on, k_off, D, v, and k_dep that are consistent with independent biochemical measurements.
In the discussion, the authors extrapolate their findings to cellular contexts. They propose that cells could fine‑tune MT architecture by regulating the expression level, processivity, and transport mode of depolymerizing motors. For instance, during mitosis, a surge in highly processive kinesin‑13 could enforce rapid MT shortening, ensuring proper spindle assembly, whereas in differentiated neurons, lower motor concentrations and more diffusive transport might preserve a broader length spectrum needed for axonal branching. The model also suggests that spatial gradients of motor concentration (e.g., near the cortex) could generate position‑dependent MT length patterns without invoking additional signaling pathways.
Finally, the paper outlines future extensions: incorporating multiple motor species with opposing activities (polymerases vs. depolymerases), accounting for MAPs that alter lattice binding affinity, and embedding the MT dynamics in a three‑dimensional cellular geometry. Such refinements would bring the theory closer to the complex reality of intracellular cytoskeletal organization.
Overall, this work provides a rigorous, experimentally validated framework that links motor protein biophysics to emergent MT length distributions, offering new insights into how cells achieve precise spatial control of their microtubule networks.
Comments & Academic Discussion
Loading comments...
Leave a Comment