Curvature and torsion in growing actin networks
Intracellular pathogens such as Listeria monocytogenes and Rickettsia rickettsii move within a host cell by polymerizing a comet-tail of actin fibers that ultimately pushes the cell forward. This dense network of cross-linked actin polymers typically exhibits a striking curvature that causes bacteria to move in gently looping paths. Theoretically, tail curvature has been linked to details of motility by considering force and torque balances from a finite number of polymerizing filaments. Here we track beads coated with a prokaryotic activator of actin polymerization in three dimensions to directly quantify the curvature and torsion of bead motility paths. We find that bead paths are more likely to have low rather than high curvature at any given time. Furthermore, path curvature changes very slowly in time, with an autocorrelation decay time of 200 seconds. Paths with a small radius of curvature, therefore, remain so for an extended period resulting in loops when confined to two dimensions. When allowed to explore a 3D space, path loops are less evident. Finally, we quantify the torsion in the bead paths and show that beads do not exhibit a significant left- or right-handed bias to their motion in 3D. These results suggest that paths of actin-propelled objects may be attributed to slow changes in curvature rather than a fixed torque.
💡 Research Summary
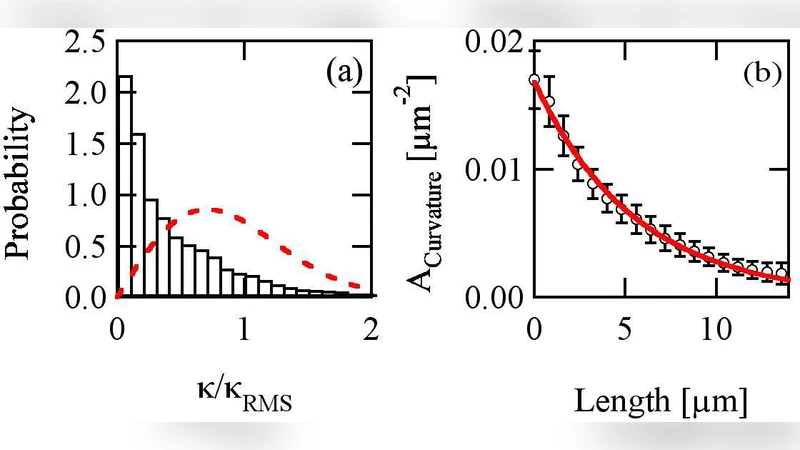
The paper investigates how actin‑driven objects move by directly measuring the curvature and torsion of their three‑dimensional trajectories. Using 3‑D fluorescence microscopy, the authors tracked silica beads coated with the prokaryotic actin‑polymerization activator ActA as they were propelled by a dense, cross‑linked actin comet tail inside living host cells. From the bead position data they computed the instantaneous curvature κ(t) – a scalar describing how sharply the path bends – and the torsion τ(t) – a measure of out‑of‑plane twisting. Statistical analysis revealed several key features. First, the curvature distribution is heavily weighted toward low values; beads are more likely to follow gently curving paths than highly curved ones. Second, the curvature autocorrelation decays exponentially with a characteristic time of roughly 200 seconds, indicating that once a particular curvature is established it persists for many minutes before slowly changing. This slow evolution explains why, when the beads are confined to a two‑dimensional plane (e.g., on a glass slide), they often trace tight loops: a small radius of curvature, once adopted, is maintained long enough to complete several revolutions. In contrast, in an unrestricted three‑dimensional environment the same slow curvature changes produce far fewer visible loops because the trajectory can wander out of the plane. Third, the measured torsion averages near zero and shows no statistically significant left‑ or right‑handed bias, implying that the actin network does not generate a persistent chiral torque on the bead. The torsion autocorrelation time (~50 s) is considerably shorter than that of curvature, suggesting that out‑of‑plane twisting fluctuates more rapidly and does not dominate the overall path shape.
These observations challenge traditional theoretical models that attribute path curvature to a fixed torque generated by a small number of polymerizing filaments. Instead, the authors propose that the dominant factor is a slowly varying curvature driven by dynamic remodeling of the actin meshwork – changes in cross‑link density, filament orientation, and polymerization rates that accumulate over time. Consequently, predictive models of actin‑propelled motility should incorporate stochastic, time‑dependent curvature rather than assume a constant torque balance. The study not only clarifies why intracellular pathogens such as Listeria and Rickettsia often follow looping trajectories but also provides a quantitative framework that could inform the design of synthetic nanomotors that harness actin polymerization for directed motion. Future work is suggested to dissect how specific actin‑binding proteins, filament nucleation rates, and external mechanical constraints modulate the observed curvature and torsion dynamics.
Comments & Academic Discussion
Loading comments...
Leave a Comment