Hidden Structure in Protein Energy Landscapes
Inherent structure theory is used to discover strong connections between simple characteristics of protein structure and the energy landscape of a Go model. The potential energies and vibrational free energies of inherent structures are highly correlated, and both reflect simple measures of networks of native contacts. These connections have important consequences for models of protein dynamics and thermodynamics.
💡 Research Summary
The paper applies inherent‑structure theory to a Go‑type protein model in order to uncover simple structural descriptors that govern the shape of the protein’s energy landscape. By performing extensive molecular‑dynamics simulations and quenching each trajectory snapshot to its nearest local minimum, the authors generated a large ensemble of inherent structures. For each inherent structure they computed two thermodynamic quantities: the potential energy (E_pot) and the vibrational free energy (F_vib) obtained from a harmonic normal‑mode analysis.
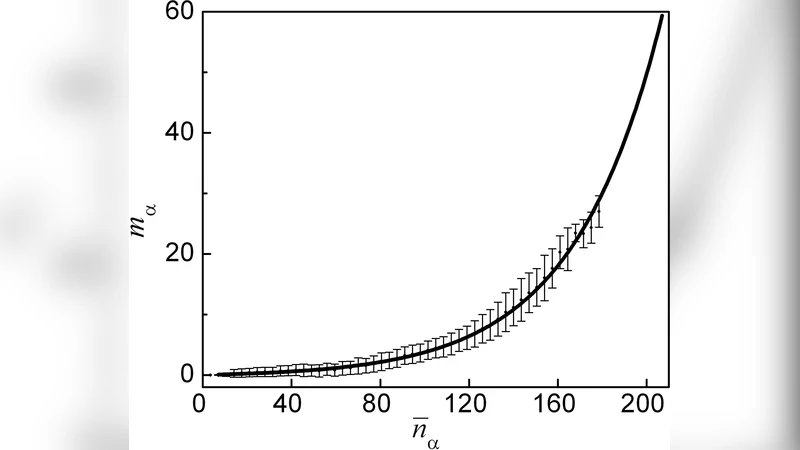
A striking linear correlation emerged between E_pot and F_vib (R² ≈ 0.94), indicating that the depth of a basin in the landscape is tightly linked to the entropy associated with its vibrational modes. To explain this correlation, the authors introduced two network‑based metrics derived from the native contact map: (1) the total number of native contacts (C) and (2) the clustering coefficient (Q), which measures the prevalence of triangular contact motifs. Both C and Q correlate strongly with the energetic quantities (C ↔ E_pot, Q ↔ F_vib), with regression slopes that are nearly identical across the whole dataset. In particular, high Q values correspond to basins that are both low in potential energy and low in vibrational entropy, reflecting a densely interconnected contact network.
The authors then examined transition pathways between inherent structures by constructing a network where nodes are minima and edges represent low‑energy barriers identified from the simulation data. They found that pathways crossing low barriers almost invariably pass through regions of the contact network with high clustering and high centrality. This suggests that a small “core” of highly interconnected contacts forms early in the folding process and acts as a kinetic bottleneck that controls the overall folding rate.
Building on these observations, the paper proposes two practical implications for protein‑folding modeling. First, Markov state models (MSMs) and Bayesian kinetic frameworks can be enriched by incorporating C, Q, and node centrality as state‑specific parameters, thereby improving the accuracy of transition‑rate estimates without dramatically increasing computational cost. Second, because the inherent‑structure ensemble captures the essential topography of the free‑energy surface, one can reconstruct a coarse‑grained landscape using only a handful of network descriptors, bypassing the need for exhaustive high‑dimensional free‑energy calculations.
In summary, the study demonstrates that the seemingly complex energy landscape of a protein can be reduced to a few simple, physically interpretable network measures of native contacts. This reduction not only clarifies the relationship between structure, energy, and dynamics but also offers a scalable route to more efficient and predictive simulations of protein folding and conformational change. Future work is suggested to test the generality of these findings on realistic all‑atom force fields and to integrate experimental observables (e.g., Φ‑value analysis) for further validation.
Comments & Academic Discussion
Loading comments...
Leave a Comment