The Interplay between Chemistry and Mechanics in the Transduction of a Mechanical Signal into a Biochemical Function
There are many processes in biology in which mechanical forces are generated. Force-bearing networks can transduce locally developed mechanical signals very extensively over different parts of the cell or tissues. In this article we conduct an overview of this kind of mechanical transduction, focusing in particular on the multiple layers of complexity displayed by the mechanisms that control and trigger the conversion of a mechanical signal into a biochemical function. Single molecule methodologies, through their capability to introduce the force in studies of biological processes in which mechanical stresses are developed, are unveiling subtle intertwining mechanisms between chemistry and mechanics and in particular are revealing how chemistry can control mechanics. The possibility that chemistry interplays with mechanics should be always considered in biochemical studies.
💡 Research Summary
Biological systems are constantly subjected to mechanical forces generated both internally and externally, ranging from muscle contraction and cell migration to tissue morphogenesis. These forces are not merely physical perturbations; they are transduced through intricate networks that convert mechanical cues into biochemical responses. The paper provides a comprehensive overview of this mechanotransduction, emphasizing the multilayered complexity that governs the conversion of a mechanical signal into a biochemical function.
At the first layer, the authors describe how force propagates through structural components such as the cytoskeleton, adhesion complexes, and the nuclear envelope. These elements possess elastic, viscoelastic, and often nonlinear mechanical properties, allowing them to channel stress to specific loci. Stress concentration creates “force hotspots” where molecular conformations are dramatically altered.
The second layer focuses on the chemical consequences of these conformational changes. When force exposes previously hidden binding sites or disrupts existing interactions, enzymatic activity can be either enhanced or inhibited. This “force‑induced activation/inhibition” provides cells with a rapid means to translate external mechanical stimuli into precise biochemical outputs, effectively linking mechanical deformation to signaling cascades.
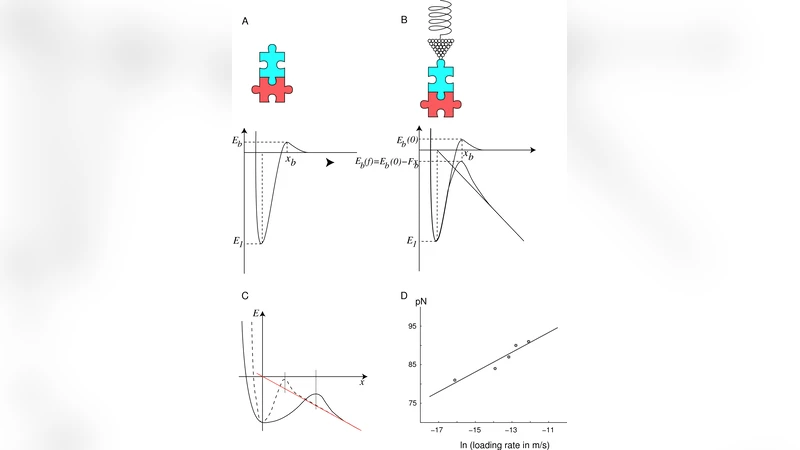
The third layer leverages single‑molecule techniques—optical tweezers, magnetic tweezers, atomic force microscopy, and related force‑spectroscopy methods—to directly apply calibrated forces to individual proteins or nucleic‑acid complexes while simultaneously monitoring chemical reactions in real time. The authors present data showing that applying a defined pico‑ to nanonewton force lowers activation energy barriers, accelerates reaction rates, and shifts reaction pathways in a force‑dependent manner. By quantifying force‑dependent reaction coordinates, they integrate mechanistic insights with classical chemical kinetics, establishing a quantitative framework for force‑chemistry coupling.
Importantly, the paper also highlights the reverse relationship: chemical modifications can modulate mechanical properties. Post‑translational modifications such as phosphorylation, methylation, or disulfide bond formation alter protein stiffness, elasticity, and stress‑propagation pathways. Consequently, the same external force can elicit different biochemical responses depending on the protein’s chemical state, creating a feedback loop where chemistry tunes mechanics and mechanics tunes chemistry.
The authors extend these concepts to tissue‑level phenomena. In cardiac muscle, rhythmic contraction synchronizes electrical signaling and metabolic pathways through coordinated mechanochemical coupling. In cancer metastasis, alterations in cellular stiffness and the expression of mechanosensitive transcription factors cooperate to facilitate invasion into new microenvironments. These examples illustrate how the multilayered mechanochemical network scales from single molecules to whole organs.
In conclusion, the paper redefines mechanotransduction as a bidirectional, multilayered process where mechanical forces and chemical reactions are interwoven. Force‑induced structural changes expose new reactive sites, while chemical modifications reshape mechanical responsiveness, establishing a dynamic feedback system. Recognizing this interplay is essential for designing experiments in cell biology, developing mechanobiology‑based therapeutics, and engineering biomaterials that harness or modulate mechanochemical signaling.
Comments & Academic Discussion
Loading comments...
Leave a Comment