Effects of the DNA state fluctuation on single-cell dynamics of self-regulating gene
A dynamical mean-field theory is developed to analyze stochastic single-cell dynamics of gene expression. By explicitly taking account of nonequilibrium and nonadiabatic features of the DNA state fluctuation, two-time correlation functions and response functions of single-cell dynamics are derived. The method is applied to a self-regulating gene to predict a rich variety of dynamical phenomena such as anomalous increase of relaxation time and oscillatory decay of correlations. Effective “temperature” defined as the ratio of the correlation to the response in the protein number is small when the DNA state change is frequent, while it grows large when the DNA state change is infrequent, indicating the strong enhancement of noise in the latter case.
💡 Research Summary
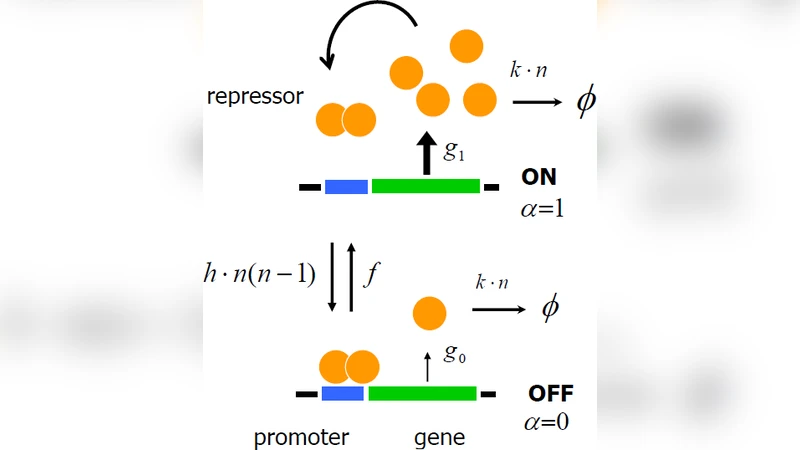
The paper develops a dynamical mean‑field theory (DMFT) that explicitly incorporates stochastic switching of a gene’s promoter between active (ON) and repressed (OFF) states. Unlike conventional approaches that assume rapid promoter equilibration and replace the discrete DNA state by a static average, this framework treats the promoter transition rate (ω) as a dynamical variable that can be comparable to or slower than protein synthesis and degradation (γ). The authors focus on a self‑regulating gene, where the protein product feeds back to repress its own transcription, providing a minimal yet biologically relevant system to explore non‑equilibrium effects.
Starting from the full master equation for the joint probability P(n,s,t) of having n protein molecules and promoter state s, the authors perform a Kramers‑Moyal expansion and close the hierarchy at second order, yielding coupled equations for the first and second moments. This procedure preserves the non‑adiabatic, non‑equilibrium contributions that would be lost in a simple adiabatic elimination of the DNA variable. Within the DMFT, they derive exact expressions for two‑time correlation functions C(t)=⟨δn(t)δn(0)⟩ and linear response functions R(t)=δ⟨n(t)⟩/δh(0), where h(t) is an external perturbation of the transcription rate. These quantities allow a direct test of the fluctuation‑dissipation relation (FDR) in a biologically realistic setting.
The analysis reveals two distinct dynamical regimes. In the fast‑switching limit (ω≫γ), promoter fluctuations are effectively averaged out; C(t) decays mono‑exponentially with a single relaxation time τ≈1/γ, and R(t) mirrors this decay. The ratio C(0)/R(0), defined as an effective temperature T_eff, remains low, indicating that noise is only modestly amplified relative to the deterministic response. Conversely, in the slow‑switching limit (ω≪γ), the promoter remains locked in a given state for long periods, causing bursts of protein production followed by extended repression. Here C(t) exhibits damped oscillations rather than simple exponential decay, reflecting the interplay between the intrinsic feedback loop and the discrete promoter dynamics. The response function becomes non‑monotonic, and T_eff grows dramatically, signifying a strong violation of the conventional FDR and an enhanced stochasticity.
A particularly striking prediction is the non‑monotonic dependence of the relaxation time τ_relax on ω. As ω decreases from the fast‑switching regime, τ_relax first increases, reaches a maximum at an intermediate ω, and then decreases again as the system enters the deep slow‑switching regime where the dynamics are dominated by rare promoter flips. This “critical‑like” slowdown suggests that cells could tune promoter kinetics to balance responsiveness against noise robustness.
The authors also discuss experimental implications. Single‑cell fluorescence time‑series or microfluidic tracking could measure C(t) and R(t) directly, allowing verification of the predicted oscillatory decay and the ω‑dependent effective temperature. Moreover, the framework can be extended to more complex networks, multi‑state promoters, or chromatin remodeling processes, offering a versatile tool for quantitative systems biology.
In summary, the paper provides a rigorous statistical‑mechanical treatment of DNA state fluctuations in gene expression, demonstrates how non‑adiabatic promoter dynamics generate rich temporal behavior—including anomalous relaxation, oscillatory correlations, and temperature‑like noise amplification—and establishes a clear connection between microscopic promoter kinetics and macroscopic cellular phenotypes. This work bridges the gap between abstract stochastic theory and experimentally observable single‑cell dynamics, opening avenues for both theoretical exploration and practical measurement of non‑equilibrium noise in genetic circuits.
Comments & Academic Discussion
Loading comments...
Leave a Comment