On Gene Duplication Models for Evolving Regulatory Networks
Background: Duplication of genes is important for evolution of molecular networks. Many authors have therefore considered gene duplication as a driving force in shaping the topology of molecular networks. In particular it has been noted that growth via duplication would act as an implicit way of preferential attachment, and thereby provide the observed broad degree distributions of molecular networks. Results: We extend current models of gene duplication and rewiring by including directions and the fact that molecular networks are not a result of unidirectional growth. We introduce upstream sites and downstream shapes to quantify potential links during duplication and rewiring. We find that this in itself generates the observed scaling of transcription factors for genome sites in procaryotes. The dynamical model can generate a scale-free degree distribution, p(k)∝ 1/k^γ, with exponent γ=1 in the non-growing case, and with γ>1 when the network is growing. Conclusions: We find that duplication of genes followed by substantial recombination of upstream regions could generate main features of genetic regulatory networks. Our steady state degree distribution is however to broad to be consistent with data, thereby suggesting that selective pruning acts as a main additional constraint on duplicated genes. Our analysis shows that gene duplication can only be a main cause for the observed broad degree distributions, if there is also substantial recombinations between upstream regions of genes.
💡 Research Summary
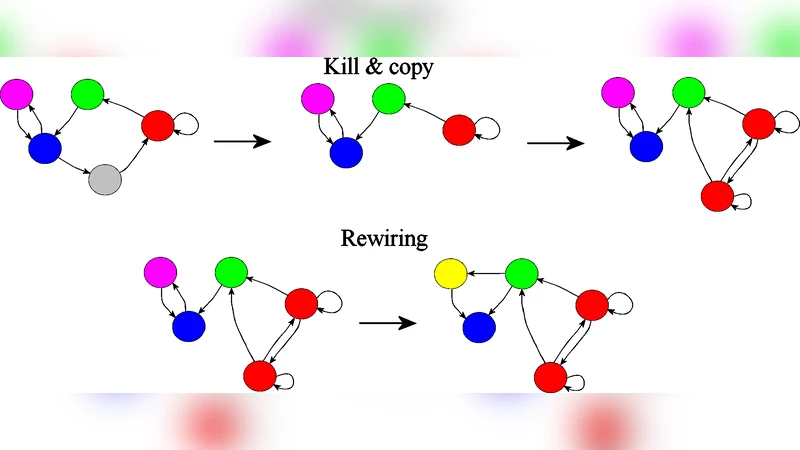
The paper investigates how gene duplication, combined with extensive recombination of upstream regulatory regions, can shape the topology of transcriptional regulatory networks. Building on earlier duplication‑rewiring models, the authors introduce a directed network framework that explicitly distinguishes between an upstream “site” (the regulatory input region) and a downstream “shape” (the output, i.e., the transcription factor protein). In each duplication event the downstream shape is copied faithfully, while the upstream site may be reshuffled with a probability p_recombination, either swapping with another gene’s site or being replaced by a novel configuration. Connections are formed when an upstream site matches a downstream shape, yielding a directed edge from regulator to target.
Two dynamical regimes are examined. In the steady‑state (non‑growing) case, duplication and recombination occur at equal rates while the total number of genes remains constant. Solving the master equation under mean‑field assumptions gives a degree distribution p(k) ∝ k⁻¹, i.e., a power‑law exponent γ≈1. In the growing regime, new genes are continuously added; the average degree rises over time and the resulting distribution follows p(k) ∝ k⁻ᵞ with γ > 1. Importantly, when the recombination probability is high, the model reproduces the empirically observed scaling of transcription‑factor (TF) numbers with genome size in prokaryotes: the number of TFs grows roughly as N^α with α≈0.5, matching data from bacteria such as E. coli.
The authors compare simulated networks to real bacterial regulatory maps. While the model captures the TF‑genome scaling, its predicted degree exponent (γ≈1 in the static case) is substantially lower than the values reported for actual networks (γ≈1.5–2.5). This discrepancy indicates that duplication plus recombination alone generate a degree distribution that is too broad. To reconcile the model with observations, the paper proposes an additional “selective pruning” process: after duplication, edges that are redundant, deleterious, or energetically costly are preferentially removed by natural selection. Introducing a pruning probability tunes the tail of the degree distribution, allowing the simulated γ to approach empirical values.
The discussion emphasizes that gene duplication can be a primary driver of the broad, scale‑free-like degree distributions seen in regulatory networks, but only when accompanied by substantial upstream recombination and subsequent selective pressure that trims excess connections. The steady‑state solution demonstrates that even without net growth, the interplay of duplication and recombination yields a power‑law, albeit with an exponent that is too low for real data. Growth amplifies the exponent, yet still falls short without pruning. The authors suggest future extensions that incorporate realistic binding‑site evolution, transcriptional repression mechanisms, and environment‑dependent selection to achieve a more faithful representation of network evolution.
In conclusion, the study provides a mathematically tractable, biologically motivated model that links gene duplication, regulatory region shuffling, and selective edge removal to the emergence of scale‑free properties and TF‑genome scaling in prokaryotic transcriptional networks. It highlights that duplication alone is insufficient; substantial recombination of upstream regions and evolutionary pruning are essential complementary forces shaping the observed network architecture.
Comments & Academic Discussion
Loading comments...
Leave a Comment