Amplification and detection of single molecule conformational fluctuation through a protein interaction network with bimodal distributions
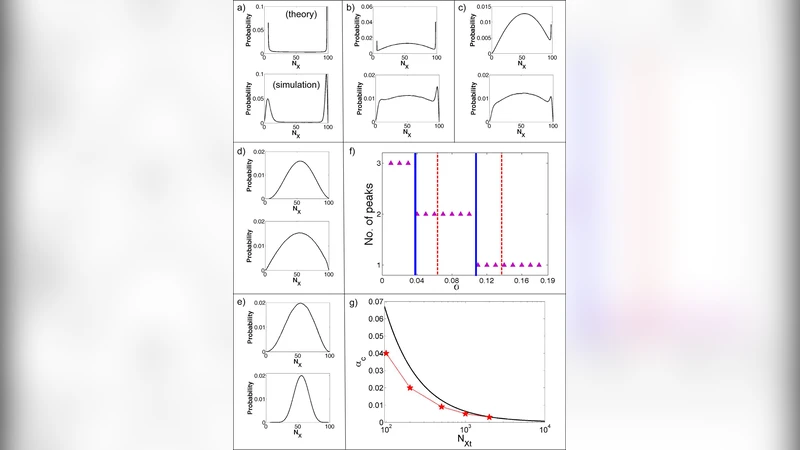
A protein undergoes conformational dynamics with multiple time scales, which results in fluctuating enzyme activities. Recent studies in single molecule enzymology have observe this “age-old” dynamic disorder phenomenon directly. However, the single molecule technique has its limitation. To be able to observe this molecular effect with real biochemical functions {\it in situ}, we propose to couple the fluctuations in enzymatic activity to noise propagations in small protein interaction networks such as zeroth order ultra-sensitive phosphorylation-dephosphorylation cycle. We showed that enzyme fluctuations could indeed be amplified by orders of magnitude into fluctuations in the level of substrate phosphorylation | a quantity widely interested in cellular biology. Enzyme conformational fluctuations sufficiently slower than the catalytic reaction turn over rate result in a bimodal concentration distribution of the phosphorylated substrate. In return, this network amplified single enzyme fluctuation can be used as a novel biochemical “reporter” for measuring single enzyme conformational fluctuation rates.
💡 Research Summary
**
The paper addresses a fundamental challenge in modern biochemistry: how to observe and quantify the conformational fluctuations of a single enzyme within its native cellular environment. While single‑molecule techniques have directly visualized “dynamic disorder” – the stochastic switching of an enzyme between distinct conformational states that possess different catalytic rates – these methods are limited by technical complexity, low throughput, and the need to isolate the molecule from its physiological context. The authors propose a clever alternative: couple the fluctuating activity of a single enzyme to a small, well‑characterized protein interaction network that acts as an amplifier, thereby converting microscopic enzyme dynamics into a macroscopic, easily measurable biochemical read‑out.
The chosen network is a zero‑order ultrasensitive phosphorylation‑dephosphorylation cycle. In this motif, a kinase (K) phosphorylates a substrate S to produce S∗, while a phosphatase (P) dephosphorylates S∗ back to S. When both enzymes operate near saturation (the zero‑order regime), the system exhibits an ultrasensitive response: a modest change in the effective activity of K or P produces a large shift in the steady‑state level of S∗. This property is analogous to a biochemical switch and is well documented in classic studies of signal transduction (Goldbeter & Koshland, 1981).
Mathematically, the authors model the enzyme of interest (E) as a two‑state Markov process: E₁ ↔ E₂ with transition rates ω₁₂ and ω₂₁. Each state has its own catalytic turnover number, k₁ and k₂, respectively. The phosphorylation cycle is described by Michaelis–Menten kinetics under the zero‑order assumption, leading to a set of coupled ordinary differential equations for
Comments & Academic Discussion
Loading comments...
Leave a Comment