Ab initio computational modeling of tumor spheroids
This paper is a review of the VBL project, where we develop and test a numerical simulator of tumor spheroids.
💡 Research Summary
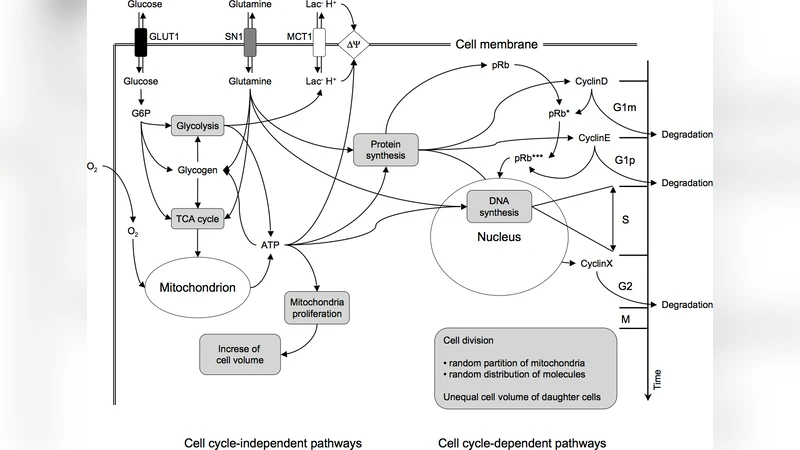
The paper provides a comprehensive review of the Virtual Biology Lab (VBL) project, whose central aim is to construct an ab‑initio computational model of tumor spheroids that relies solely on first‑principles physics and biology. The authors begin by outlining the shortcomings of conventional tumor modeling approaches, which often depend on empirical fitting, limited experimental data, and coarse‑grained assumptions that obscure mechanistic insight. In response, the VBL framework treats each cell as an autonomous agent governed by a set of ordinary differential equations describing metabolism (glycolysis, oxidative phosphorylation), cell‑cycle progression, and death pathways (apoptosis, necrosis). Surrounding the cellular lattice is a continuous microenvironment modeled by diffusion‑reaction partial differential equations for oxygen, nutrients, and therapeutic agents. Mechanical interactions are captured through a non‑linear elastic model that accounts for cell‑cell adhesion, compressibility, and the emergent pressure field within the spheroid.
A hybrid numerical scheme is employed: a lattice‑free particle method handles the discrete cell dynamics, while a finite‑volume discretization solves the continuum fields. This combination avoids the overhead of generating and updating a spatial grid for the cells while preserving high fidelity in the transport of solutes and mechanical stresses. The implementation is written in C++ with CUDA extensions, enabling GPU acceleration that reduces runtime dramatically for spheroids containing up to several hundred thousand cells. Parameter estimation is performed using Bayesian optimization and Markov‑Chain Monte Carlo sampling, allowing the model to be calibrated against experimental measurements of spheroid growth curves, internal oxygen gradients, and drug penetration profiles.
Validation against in‑vitro data demonstrates that the model accurately reproduces the characteristic sigmoidal growth of spheroids, the abrupt expansion of hypoxic cores once the diameter exceeds ~400 µm, and the spatially heterogeneous cell‑death patterns observed after drug administration. Sensitivity analyses reveal that oxygen diffusion coefficients and cellular elastic moduli are the dominant determinants of spheroid size and internal stress distribution, respectively. These findings suggest that the model can be used to predict how alterations in microenvironmental parameters might influence therapeutic outcomes.
The discussion acknowledges several limitations. Intracellular signaling networks are heavily simplified, precluding a detailed representation of genetic heterogeneity and adaptive responses. Angiogenesis, immune cell infiltration, and extracellular matrix remodeling are not yet incorporated, which restricts the model’s applicability to early‑stage avascular tumors. Moreover, scaling the simulation to spheroids larger than one million cells remains computationally demanding despite GPU acceleration.
Future directions outlined by the authors include extending the framework to a true multi‑scale platform that integrates genome‑scale metabolic models, stochastic gene expression, and immune‑tumor interactions. They propose leveraging distributed computing clusters and advanced load‑balancing techniques to overcome current scalability bottlenecks. Finally, the authors envision coupling the VBL simulator with patient‑specific imaging and omics data to create a personalized predictive tool for optimizing chemotherapy and radiotherapy regimens. In summary, the VBL project represents a significant step toward mechanistically grounded, high‑resolution virtual tumor models, offering both a testbed for hypothesis generation and a potential foundation for precision oncology applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment