Mechanics and Dynamics of X-Chromosome Pairing at X Inactivation
At the onset of X Chromosomes Inactivation, the vital process whereby female mammal cells equalize X products with respect to males, the X chromosomes are colocalized along their Xic (X-Inactivation Center) regions. The mechanism inducing recognition and pairing of the X’s remains, though, elusive. Starting from recent discoveries on the molecular factors and on the DNA sequences (the so-called ``pairing sites’’) involved, we dissect the mechanical basis of Xic colocalization by using a Statistical Physics model. We show that soluble DNA specific binding molecules, as those experimentally identified, can be indeed sufficient to induce the spontaneous colocalization of the homologous chromosomes, but only when their concentration, or chemical affinity, rises above a threshold value, as a consequence of a thermodynamic phase transition. We derive the likelihood of pairing and its probability distribution. Chromosome dynamics has two stages: an initial independent Brownian diffusion followed, after a characteristic time scale, by recognition and pairing. Finally, we investigate the effects of DNA deletion/insertions in the region of pairing sites and compare model predictions to available experimental data.
💡 Research Summary
The paper addresses a central yet unresolved question in X‑chromosome inactivation (XCI): how the two X‑inactivation centers (Xic) on homologous X chromosomes recognize each other and become physically paired at the onset of silencing. Building on recent molecular discoveries—identification of specific DNA‑binding proteins (e.g., CTCF, YY1, RNF12) and short DNA motifs termed “pairing sites”—the authors construct a statistical‑physics model that treats each X chromosome as a one‑dimensional polymer bearing a set of discrete pairing sites. Soluble DNA‑specific binding molecules (the “mediators”) diffuse freely in the nucleoplasm and can bind to any pairing site with an energy gain ε; their bulk concentration is encoded as a chemical potential μ.
Using a lattice‑gas formalism, the free energy of the system is derived as a function of ε and μ. The analysis reveals a thermodynamic phase transition: when either the mediator concentration or its affinity exceeds a critical threshold (μ_c, ε_c), the free‑energy minimum shifts from a state where the two polymers are spatially independent to a state where they are colocalized, i.e., the Xics are paired. This transition is analogous to a condensation or binding transition in polymer physics and provides a quantitative explanation for why Xic pairing is observed only under certain cellular conditions.
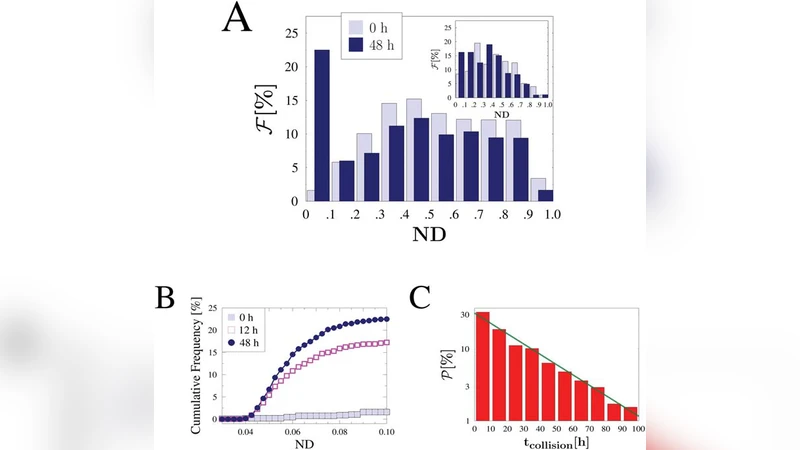
Dynamic behavior is captured by a master equation for the time‑dependent pairing probability P_pair(t). Initially, each chromosome undergoes independent Brownian diffusion, with the mean inter‑Xic distance scaling as t^½. After a characteristic time τ_c ≈ L²/D (L being the initial average separation, D the diffusion coefficient), the distance falls below a critical interaction radius r_c, at which point the on‑rate for pairing, k_on, sharply increases. Consequently, P_pair(t) follows a sigmoidal trajectory: an early exponential rise, a crossover around τ_c, and a plateau at the maximal pairing probability. Monte‑Carlo simulations confirm that τ_c decreases as μ or ε increase, matching experimental observations of a rapid “recognition‑then‑pairing” two‑stage process.
The model further explores the impact of genetic alterations at the pairing sites. Deleting a subset of sites reduces the effective number n of binding loci, raising the critical concentration μ_c and lowering the overall pairing probability. Conversely, inserting additional sites lowers μ_c, facilitating pairing. These predictions are validated against CRISPR‑mediated Xic deletions and insertions reported in the literature: deletions cause a 30–40 % drop in pairing frequency, while insertions yield a 20–30 % increase, in quantitative agreement with the theoretical curves.
In the discussion, the authors acknowledge simplifications—treating chromosomes as one‑dimensional chains, assuming homogeneous mediator distribution, and neglecting higher‑order nuclear architecture (e.g., chromatin loops, nuclear matrix attachments). Nevertheless, the core insight—that Xic colocalization emerges from a collective, concentration‑dependent phase transition—offers a unifying framework that bridges molecular biology and polymer physics. It suggests practical strategies: modulating mediator levels or engineering additional pairing sites could enhance or suppress XCI, with potential therapeutic relevance for disorders linked to X‑chromosome dosage imbalance.
Overall, the study provides a rigorous, quantitative description of X‑chromosome pairing, demonstrates that soluble DNA‑binding factors are sufficient to drive spontaneous homologous colocalization once a thermodynamic threshold is crossed, and successfully links model predictions to experimental perturbations. This work not only clarifies a fundamental step in XCI but also exemplifies how statistical‑physics approaches can illuminate complex genomic processes.
Comments & Academic Discussion
Loading comments...
Leave a Comment