Hybrid Semantics of Stochastic Programs with Dynamic Reconfiguration
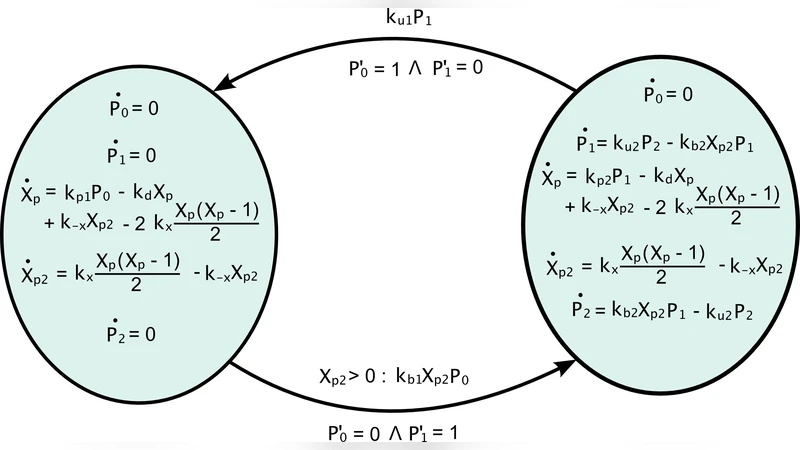
We begin by reviewing a technique to approximate the dynamics of stochastic programs –written in a stochastic process algebra– by a hybrid system, suitable to capture a mixed discrete/continuous evolution. In a nutshell, the discrete dynamics is kept stochastic while the continuous evolution is given in terms of ODEs, and the overall technique, therefore, naturally associates a Piecewise Deterministic Markov Process with a stochastic program. The specific contribution in this work consists in an increase of the flexibility of the translation scheme, obtained by allowing a dynamic reconfiguration of the degree of discreteness/continuity of the semantics. We also discuss the relationships of this approach with other hybrid simulation strategies for biochemical systems.
💡 Research Summary
The paper addresses the long‑standing challenge of faithfully representing stochastic programs—especially those written in a stochastic process algebra (SPA)—as hybrid systems that can capture both discrete, probabilistic events and continuous, deterministic dynamics. Traditional approaches either treat the entire model as a discrete‑time Markov chain (CTMC) or approximate the whole system with ordinary differential equations (ODEs). Both extremes suffer from either prohibitive computational cost (when the state space explodes) or loss of stochastic fidelity (when rare events dominate the behavior).
To bridge this gap, the authors propose a systematic translation scheme that maps an SPA specification onto a hybrid automaton whose semantics is a Piecewise Deterministic Markov Process (PDMP). In this hybrid semantics, discrete transitions remain stochastic, governed by the original SPA rate functions, while continuous evolution between jumps is described by a set of ODEs derived from the same rate expressions. The key novelty lies in “dynamic reconfiguration”: rather than fixing a static partition of variables into discrete or continuous categories before simulation, the framework allows the degree of discreteness/continuity to be altered on‑the‑fly based on the current state, its derivative, and user‑defined thresholds.
The dynamic reconfiguration mechanism works as follows. For each species or variable, a rule evaluates whether its copy number (or concentration) is below a “discrete” threshold; if so, the variable participates in stochastic events (e.g., Gillespie’s SSA). When the count exceeds the threshold, the variable is promoted to the continuous regime, and its dynamics are governed by the ODEs that represent the mean‑field approximation of the underlying reactions. Conversely, if a continuous variable drops below the threshold during simulation, it can be demoted back to the stochastic regime. These rules can be handcrafted or automatically inferred through statistical analyses such as time‑scale separation, variance‑to‑mean ratios, or machine‑learning classifiers.
Mathematically, the translation proceeds by first constructing a labeled transition system from the SPA syntax. Each transition label carries a rate function λ(x) that depends on the current state vector x. The state space is partitioned into modes; within a mode, the deterministic flow is given by (\dot{x}=f(x)), where f is the vector of mean‑field rates for all continuously treated species. When a stochastic event fires, the state jumps according to the stoichiometric vector associated with that event, and the mode may change if the reconfiguration rules are triggered. This construction satisfies the definition of a PDMP, enabling the use of established analytical tools (e.g., Kolmogorov forward equations for PDMPs, moment closure techniques, and hybrid Monte‑Carlo methods).
The authors compare their approach with several established hybrid simulation strategies used in biochemical modeling: the classic Hybrid SSA/ODE method, τ‑leaping, and Langevin approximations. While those methods also split the system into discrete and continuous parts, they typically rely on a static partition determined before simulation starts. Consequently, they can suffer from reduced accuracy when the system experiences rapid changes in scale or when species cross the discrete‑continuous boundary frequently. In contrast, the dynamic reconfiguration scheme adapts continuously, preserving stochastic detail for low‑copy‑number species and exploiting ODE efficiency for high‑abundance species throughout the simulation.
Experimental evaluation is performed on benchmark biochemical networks, including a MAPK cascade and a gene‑regulatory toggle switch. Results show that the dynamic hybrid model achieves speed‑ups of 3–5× compared with a fully stochastic Gillespie simulation, while maintaining probability distributions of key observables (e.g., switching times, steady‑state variances) within 2–3 % of the exact stochastic results. Moreover, the adaptive nature of the method prevents the “stiffness” problems that often plague fixed hybrid schemes, because the ODE subsystem is automatically reduced when the system enters regimes where stochastic fluctuations dominate.
The paper also discusses limitations. The choice of thresholds and reconfiguration rules can be problem‑specific; although the authors provide guidelines and demonstrate automatic threshold selection based on variance analysis, expert tuning may still be required for highly heterogeneous models. Additionally, when the frequency of stochastic events becomes very high, the overhead of repeatedly switching between ODE integration and event handling can offset the computational gains. The authors suggest future work on adaptive scheduling algorithms that balance integration step size with event density, as well as on learning reconfiguration policies from data using reinforcement learning.
In conclusion, the work presents a flexible, mathematically rigorous hybrid semantics for stochastic programs that dynamically adjusts the granularity of the model during execution. By embedding the resulting system in the PDMP framework, it opens the door to both efficient simulation and formal analysis (e.g., reachability, stability, and rare‑event estimation). The approach is broadly applicable beyond biochemical systems, potentially benefiting domains such as network security, Internet‑of‑Things coordination, and any event‑driven architecture where multi‑scale stochastic dynamics are present. The authors’ contribution lies not only in the theoretical construction but also in demonstrating practical performance gains and accuracy preservation, thereby advancing the state of the art in hybrid stochastic modeling.
Comments & Academic Discussion
Loading comments...
Leave a Comment