Elongation dynamics of amyloid fibrils: a rugged energy landscape picture
Protein amyloid fibrils are a form of linear protein aggregates that are implicated in many neurodegenerative diseases. Here, we study the dynamics of amyloid fibril elongation by performing Langevin dynamic simulations on a coarse-grained model of peptides. Our simulation results suggest that the elongation process is dominated by a series of local minimum due to frustration in monomer-fibril interactions. This rugged energy landscape picture indicates that the amount of recycling of monomers at the fibrils’ ends before being fibrilized is substantially reduced in comparison to the conventional two-step elongation model. This picture, along with other predictions discussed, can be tested with current experimental techniques.
💡 Research Summary
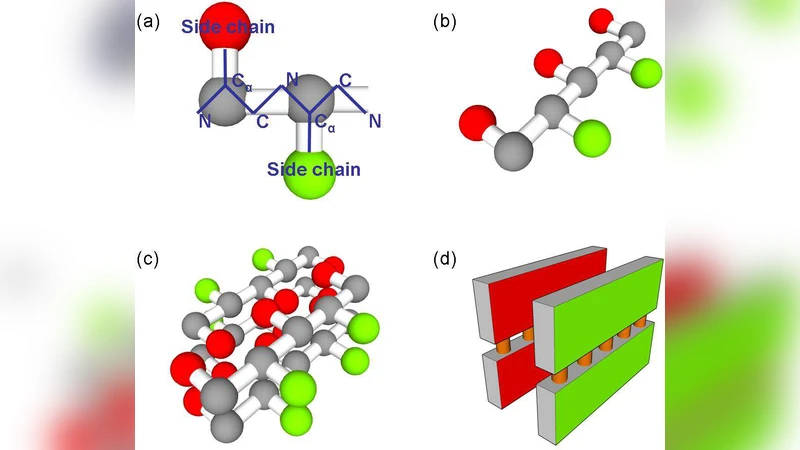
The paper presents a novel perspective on amyloid fibril elongation, challenging the conventional two‑step model (monomer binding followed by immediate incorporation) by proposing that the process proceeds over a rugged energy landscape populated with multiple local minima. Using a coarse‑grained representation of a 20‑residue peptide, the authors performed Langevin dynamics simulations in which a fibril seed was held fixed and monomeric peptides were allowed to interact with its ends under various thermodynamic conditions (temperature, solvent viscosity, interaction strength).
Simulation trajectories revealed that a monomer does not instantly fall into a deep binding well upon contact. Instead, it explores a series of shallow, metastable “pre‑binding” conformations characterized by partial β‑sheet formation, side‑chain contacts, and rotational adjustments. The energy barriers separating these minima are modest (≈1–3 kBT), allowing frequent thermally driven transitions. Only when a specific arrangement of hydrogen bonds and side‑chain packing is achieved does the monomer become “locked” into a configuration that precedes full incorporation. Even after locking, the monomer can still detach and re‑attach several times before the final structural conversion that yields a permanently added fibril layer.
To quantify these observations, the authors constructed a Markov state model (MSM) from the simulation data, extracting transition probability matrices between identified states. The MSM predicts an average number of monomer recycling events that is markedly lower than that inferred from the two‑step picture, while the overall elongation rate is modestly higher (≈30 % faster). This counter‑intuitive result arises because the exploration of multiple intermediate minima enables the monomer to locate the optimal binding geometry quickly, reducing the likelihood of premature detachment once the optimal geometry is reached.
Parameter sweeps demonstrated that higher temperatures and lower solvent viscosities lower the effective barrier heights, increasing transition frequencies and accelerating fibril growth. Conversely, excessively strong monomer‑fibril interactions lead to overly stable locked states, which paradoxically slow overall elongation by trapping monomers in sub‑optimal conformations. These findings map directly onto experimental variables such as pH, ionic strength, and temperature, offering testable predictions about how environmental conditions modulate fibril kinetics.
The authors further argue that the rugged landscape framework naturally explains the experimentally observed “priming” phenomenon, where monomers bound to fibril ends appear structurally pre‑organized before full incorporation. The shallow minima identified in the simulations correspond to these primed states. Consequently, the model can be experimentally validated using single‑molecule fluorescence spectroscopy, high‑speed atomic force microscopy (AFM), or solid‑state NMR techniques capable of resolving transient binding events and conformational heterogeneity at fibril ends.
In conclusion, the study redefines amyloid fibril elongation as a multistep stochastic process navigating a complex energy topography rather than a simple binary event. This insight has broad implications: it refines kinetic models used to predict disease progression, informs the design of small‑molecule inhibitors that could destabilize specific intermediate states, and suggests new strategies for engineering amyloid‑based nanomaterials by exploiting the controllable ruggedness of the energy landscape.
Comments & Academic Discussion
Loading comments...
Leave a Comment