Mechanisms of Size Control and Polymorphism in Viral Capsid Assembly
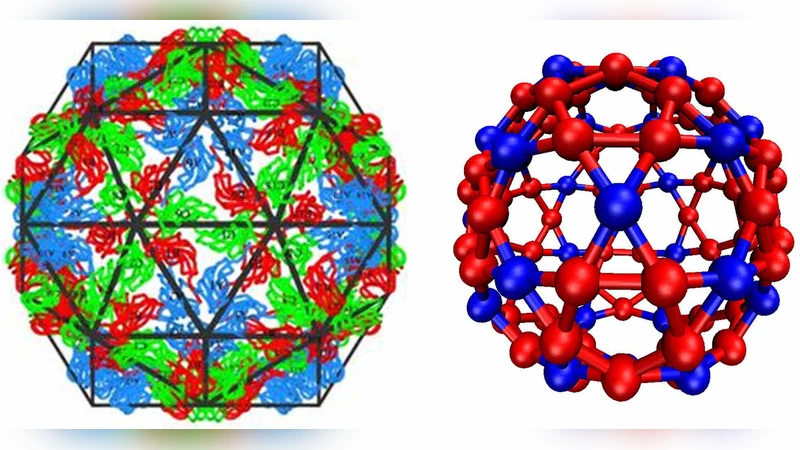
We simulate the assembly dynamics of icosahedral capsids from subunits that interconvert between different conformations (or quasi-equivalent states). The simulations identify mechanisms by which subunits form empty capsids with only one morphology but adaptively assemble into different icosahedral morphologies around nanoparticle cargoes with varying sizes, as seen in recent experiments with brome mosaic virus (BMV) capsid proteins. Adaptive cargo encapsidation requires moderate cargo-subunit interaction strengths; stronger interactions frustrate assembly by stabilizing intermediates with incommensurate curvature. We compare simulation results to experiments with cowpea chlorotic mottle virus empty capsids and BMV capsids assembled on functionalized nanoparticles and suggest new cargo encapsidation experiments. Finally, we find that both empty and templated capsids maintain the precise spatial ordering of subunit conformations seen in the crystal structure even if interactions that preserve this arrangement are favored by as little as the thermal energy, consistent with experimental observations that different subunit conformations are highly similar.
💡 Research Summary
The paper presents a comprehensive computational study of icosahedral viral capsid assembly, focusing on how capsid proteins that can interconvert among quasi‑equivalent conformations achieve size control and polymorphism. Using a coarse‑grained model, each capsid subunit is allowed to adopt three distinct conformational states, with state‑dependent interaction energies that differ by only a few kBT, reflecting the subtle energetic differences observed in crystal structures of real viruses. The authors implement two simulation frameworks: (1) Monte‑Carlo simulations of empty capsid formation across a wide range of subunit concentrations and subunit‑subunit binding strengths, and (2) Gillespie‑type kinetic simulations of capsid assembly around rigid spherical nanoparticles of varying radii, where the nanoparticle provides a templating surface that interacts with subunits via a tunable attractive potential.
Key findings from the empty‑capsid simulations show that when subunit‑subunit binding exceeds a modest threshold, the system rapidly converges to a single morphology (either T = 1 or T = 3, depending on the chosen parameters). Intermediate curvature intermediates are thermodynamically unstable and dissolve, indicating that the intrinsic curvature of the subunits dictates a unique optimal capsid size in the absence of external constraints.
In the templated‑assembly simulations, the strength of the capsid‑nanoparticle interaction emerges as the critical control knob. At intermediate interaction energies (≈ 2–3 kBT per contact), subunits can adjust their quasi‑equivalent state distribution to match the curvature imposed by the particle, leading to adaptive encapsidation: small particles (≈ 20 nm) favor T = 1 capsids, medium particles (≈ 30 nm) favor T = 3, and larger particles (≈ 40 nm) promote higher‑T structures such as T = 4. This adaptive behavior reproduces recent experimental observations with Brome Mosaic Virus (BMV) capsid proteins on functionalized nanoparticles. Conversely, when the capsid‑particle attraction is too strong (≥ 5 kBT per contact), the system becomes frustrated: the particle’s curvature cannot be reconciled with the preferred subunit curvature, leading to the stabilization of malformed intermediates containing curvature defects (e.g., misplaced five‑fold vertices). These defective intermediates stall further growth, dramatically reducing assembly yields—a phenomenon the authors term “frustration‑induced kinetic trap.”
An additional, perhaps surprising, result is that the precise spatial ordering of subunit conformations observed in high‑resolution crystal structures is maintained even when the energetic bias favoring that ordering is as small as a single thermal energy unit (kBT). This suggests that quasi‑equivalence is a robust, entropy‑driven principle that can be preserved under very weak energetic discrimination, consistent with experimental reports that different conformational states of capsid proteins are structurally almost indistinguishable.
The authors compare their simulation outcomes with experimental data on Cowpea Chlorotic Mottle Virus (CCMV) empty capsids and BMV capsids assembled on nanoparticles, finding quantitative agreement in both the preferred T‑numbers and the dependence of assembly efficiency on interaction strength. They propose several concrete experimental tests: (i) systematic variation of ligand density or length on nanoparticle surfaces to fine‑tune capsid‑particle affinity, (ii) use of cryo‑EM and single‑particle reconstruction to map the distribution of quasi‑equivalent states in assembled capsids of different sizes, and (iii) real‑time monitoring of assembly pathways using single‑molecule FRET to detect curvature‑adjustment events.
Overall, the study elucidates the dual role of intrinsic subunit curvature and extrinsic templating forces in governing viral capsid size selection and polymorphism. It provides a predictive framework for designing custom nanocontainers by engineering capsid‑protein interactions and nanoparticle templates, and it highlights the delicate balance between sufficient affinity (to drive encapsidation) and excessive affinity (which leads to kinetic traps). The work bridges theoretical modeling with experimental virology and nanotechnology, offering actionable insights for both fundamental virus assembly research and the development of virus‑based nanomaterials.
Comments & Academic Discussion
Loading comments...
Leave a Comment