A correlated topic model of Science
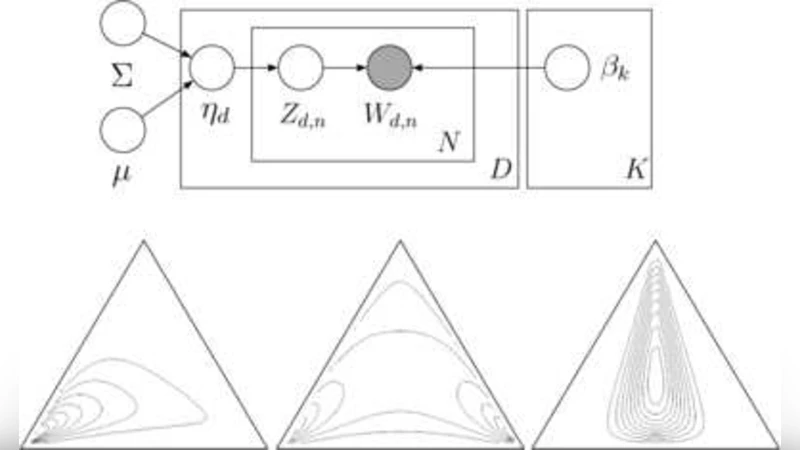
Topic models, such as latent Dirichlet allocation (LDA), can be useful tools for the statistical analysis of document collections and other discrete data. The LDA model assumes that the words of each document arise from a mixture of topics, each of which is a distribution over the vocabulary. A limitation of LDA is the inability to model topic correlation even though, for example, a document about genetics is more likely to also be about disease than X-ray astronomy. This limitation stems from the use of the Dirichlet distribution to model the variability among the topic proportions. In this paper we develop the correlated topic model (CTM), where the topic proportions exhibit correlation via the logistic normal distribution [J. Roy. Statist. Soc. Ser. B 44 (1982) 139–177]. We derive a fast variational inference algorithm for approximate posterior inference in this model, which is complicated by the fact that the logistic normal is not conjugate to the multinomial. We apply the CTM to the articles from Science published from 1990–1999, a data set that comprises 57M words. The CTM gives a better fit of the data than LDA, and we demonstrate its use as an exploratory tool of large document collections.
💡 Research Summary
The paper addresses a fundamental limitation of Latent Dirichlet Allocation (LDA), namely its inability to capture correlations among topics. LDA assumes that each document’s topic proportions are drawn from a Dirichlet distribution, which enforces near‑independence among topics. In many real‑world corpora—especially scientific literature—certain topics tend to co‑occur (e.g., genetics and disease) while others are mutually exclusive. To overcome this, the authors introduce the Correlated Topic Model (CTM), which replaces the Dirichlet prior with a logistic‑normal prior. The logistic‑normal is obtained by applying a soft‑max transformation to a multivariate Gaussian; its mean vector and covariance matrix directly encode the expected prevalence of each topic and the pairwise correlations among them. Positive off‑diagonal entries in the covariance matrix indicate that two topics are likely to appear together, while negative entries capture antagonistic relationships.
Because the logistic‑normal is not conjugate to the multinomial likelihood, exact Bayesian inference is intractable. The authors therefore develop a variational Bayes algorithm that approximates the posterior over the latent variables. The variational distribution factorizes into a Gaussian over the log‑odds of the topic proportions (parameterized by a mean μ and covariance Σ) and a multinomial over word‑topic assignments. An EM‑style coordinate ascent updates the variational parameters: the E‑step computes expected sufficient statistics given the current μ and Σ, while the M‑step maximizes a lower bound on the marginal log‑likelihood with respect to μ and Σ. To keep computation feasible for large vocabularies and many topics, they employ a Laplace approximation for the Gaussian integrals and stochastic optimization techniques that avoid forming the full K×K covariance matrix explicitly.
The model is evaluated on a massive corpus consisting of all Science journal articles published between 1990 and 1999, amounting to roughly 57 million words across about 20 000 documents. After standard preprocessing, the authors retain a vocabulary of 10 000 terms and fit both CTM and LDA with the same number of topics (K = 100). Perplexity—a standard measure of predictive performance—is consistently lower for CTM across all held‑out test sets, indicating a better fit to the data. The advantage grows as the number of topics increases, confirming that the ability to model topic correlations becomes more valuable in richer latent spaces.
Beyond quantitative metrics, the authors examine the learned covariance matrix to interpret the semantic relationships among topics. Visualizations reveal intuitive patterns: a strong positive correlation between “genetics” and “disease,” a tight coupling of “astronomy” and “physics,” and weaker negative links between “social sciences” and “molecular biology.” By projecting documents onto the latent space, they demonstrate how CTM can be used for exploratory analysis, such as tracking the rise of genomics‑related research in the mid‑1990s—a trend that aligns with the launch of the Human Genome Project.
The paper also discusses limitations and future extensions. The full covariance matrix scales quadratically with the number of topics, posing memory and computational challenges for very large K. Possible remedies include low‑rank approximations, sparsity‑inducing priors, or structured covariance models. Moreover, the static CTM does not capture temporal evolution; a dynamic version could model how topic correlations shift over time. Integrating auxiliary metadata (author affiliations, journal sections) is another promising direction.
In summary, the Correlated Topic Model provides a principled and scalable way to incorporate topic correlations into probabilistic topic modeling. It outperforms LDA on a substantial scientific corpus, yields interpretable correlation structures, and opens up new possibilities for large‑scale document exploration and trend analysis.
Comments & Academic Discussion
Loading comments...
Leave a Comment