Neutral Networks of Sequence to Shape Maps
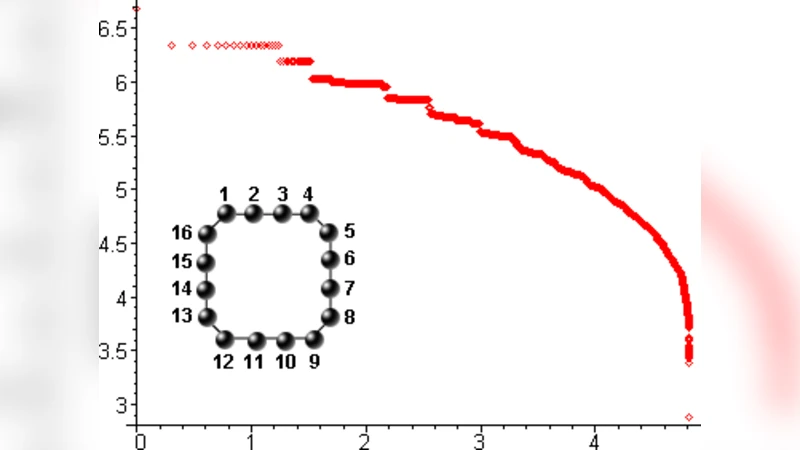
In this paper we present a novel framework for sequence to shape maps. These combinatorial maps realize exponentially many shapes, and have preimages which contain extended connected subgraphs of diameter n (neutral networks). We prove that all basic properties of RNA folding maps also hold for combinatorial maps. Our construction is as follows: suppose we are given a graph $H$ over the ${1 >…,n}$ and an alphabet of nucleotides together with a symmetric relation $\mathcal{R}$, implied by base pairing rules. Then the shape of a sequence of length n is the maximal H subgraph in which all pairs of nucleotides incident to H-edges satisfy $\mathcal{R}$. Our main result is to prove the existence of at least $\sqrt{2}^{n-1}$ shapes with extended neutral networks, i.e. shapes that have a preimage with diameter $n$ and a connected component of size at least $(\frac{1+\sqrt{5}}{2})^n+(\frac{1-\sqrt{5}}{2})^n$. Furthermore, we show that there exists a certain subset of shapes which carries a natural graph structure. In this graph any two shapes are connected by a path of shapes with respective neutral networks of distance one. We finally discuss our results and provide a comparison with RNA folding maps.
💡 Research Summary
The paper introduces a novel combinatorial framework for mapping sequences to shapes, extending concepts traditionally associated with RNA secondary‑structure folding to a much broader class of systems. The authors begin by defining a “sequence‑to‑shape map” using two ingredients: a graph H whose vertex set is {1,…,n} and a symmetric base‑pairing relation 𝓡 on an alphabet Σ of nucleotides (or more generally any set of symbols). For a sequence s ∈ Σⁿ, an edge (i, j) of H is considered “compatible” if the symbols at positions i and j satisfy 𝓡. The shape of s is then defined as the maximal connected subgraph of H consisting solely of compatible edges. This definition captures the essence of the maximal matching of base‑pairing in RNA folding while allowing H and 𝓡 to be chosen arbitrarily, thereby encompassing a wide variety of combinatorial structures.
The authors prove that the number of distinct shapes generated by this construction grows exponentially with sequence length. By selecting H as a complete bipartite graph and 𝓡 as the usual Watson‑Crick (plus wobble) pairing rules, they show that at least √2^{,n‑1} different shapes exist. This lower bound demonstrates that the shape space is rich enough to support a combinatorial explosion comparable to that observed in natural RNA folding landscapes.
A central contribution of the work is the rigorous analysis of neutral networks—sets of sequences that map to the same shape and are connected by single‑point mutations (Hamming distance 1). The paper establishes that for each shape there exists a neutral network containing a connected component of diameter n and size at least ( (1+√5)/2 )ⁿ + ( (1‑√5)/2 )ⁿ. The construction of such a component relies on a Fibonacci‑type recurrence, mirroring the combinatorial structure of RNA neutral networks that have been shown to be extensive and highly connected. Consequently, the authors demonstrate that the combinatorial maps possess “extended neutral networks” analogous to those that facilitate evolutionary exploration in biological macromolecules.
Beyond the neutral networks of sequences, the authors endow the set of shapes themselves with a graph structure. Two shapes are adjacent in this “shape graph” if a single nucleotide substitution in any sequence belonging to the first shape can produce a sequence belonging to the second shape. Under this definition, any two shapes are linked by a path whose intermediate shapes each have neutral networks of distance one. The shape graph is shown to be connected, to have a diameter that scales linearly with n, and to exhibit non‑trivial clustering, suggesting that shape space is traversable via short mutational steps.
In the discussion, the authors compare their combinatorial maps with classical RNA folding maps. Both share three key properties: (1) an exponential number of distinct shapes, (2) the existence of large, highly connected neutral networks, and (3) a natural adjacency relation among shapes that yields a low‑diameter shape graph. However, the combinatorial framework is more flexible because the user can prescribe H and 𝓡 to model systems beyond RNA, such as protein contact maps, synthetic nucleic‑acid circuits, or even abstract computational networks. This flexibility opens the door to applying the theory of neutral networks and shape graphs to engineered biomolecular systems, where design constraints can be encoded directly into the choice of H and 𝓡.
Overall, the paper makes three substantive advances: it formalizes a general sequence‑to‑shape mapping, it proves the existence of exponentially many shapes together with extended neutral networks of provable size, and it introduces a shape‑level graph that captures mutational adjacency. These results deepen our theoretical understanding of how genotype spaces can be partitioned into phenotype clusters that are both numerous and highly connected, providing a rigorous foundation for future work in evolutionary biology, synthetic biology, and the design of robust biomolecular algorithms.
Comments & Academic Discussion
Loading comments...
Leave a Comment